EXPERIMENTS
Here, you will find all the necessary information about the wetlab materials and methods we used in the course of our project.
Escherichia coli vectors
pET28
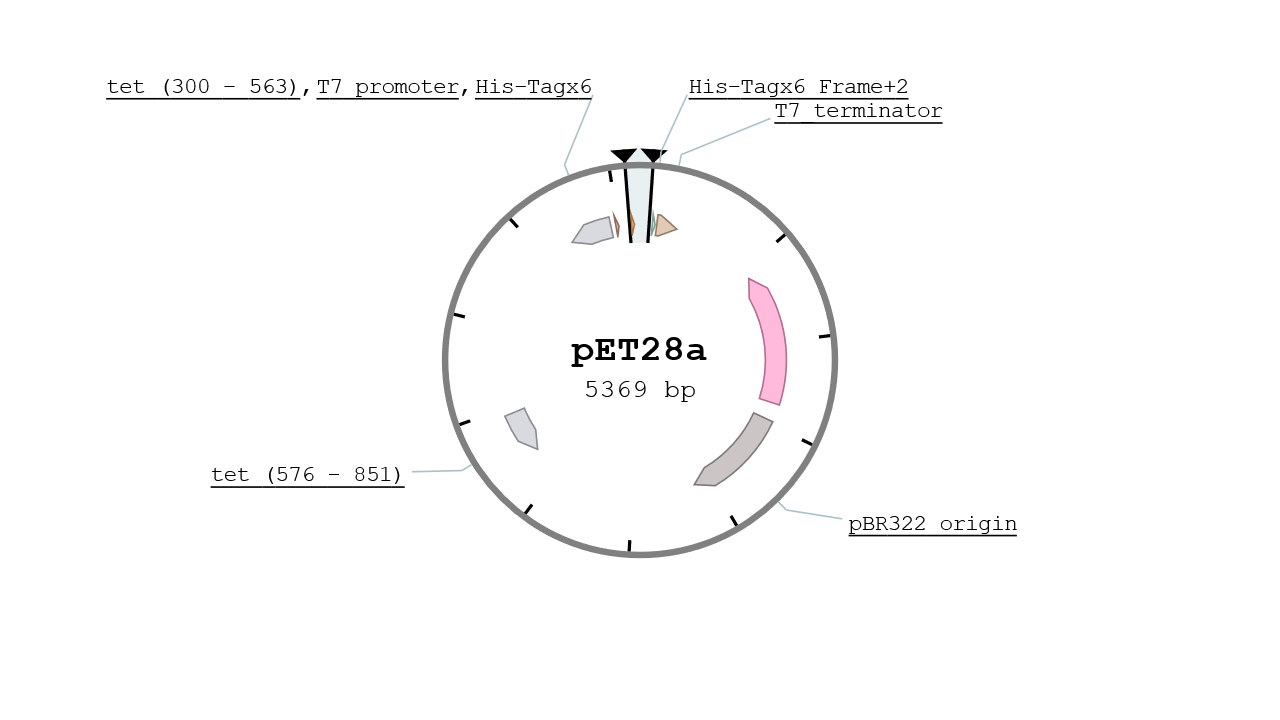
The pET28 vector contains two His Tags in its MCS and is specified for protein expression at high yield. It gives the strain a resistance to Kanamycin and its replication origin is the pBR322 which is a mid - low copy number plasmid (~10). The vector used was a kind gift from IBCG library.
pETDuet-1
The pETDuet-1 is, like the pET28, specialized in high yield protein production. Its main difference is that it contains two MCS and can therefore express two proteins at the same time under the same expression conditions. It bears ampicilin resistance and is a mid - low copy number plasmid. The plasmid used was a gift from IBCG library.

Pichia pastoris vectors
pPICZ alpha
The pPICZalpha is an integrative plasmid for Pichia pastoris protein expression and suitable for E. coli replication. Its MCS is suitable for protein expession using the methanol inductible AOX1 Pomoter (PAOX1) and offering the alpha factor secretion signal for protein purification from culture supernatant. It contains a His tag and a Myc Tag for affinity purification stategies.

pGAPZalpha
The pGAPZalpha is roughly the same as the pPICZalpha with a different promoter that is the GAP Promoter, a glucose inductible promoter.

Primer used
- Cerberus Forward : TAAGAAGGAGATATACCATGGCGGAAGCGGGTATCACC
- Cerberus Reverse : CTCGAGTGCGGCCGCAAGCTTCGGATCGTCCTATGATGGAGG
- Sirius Forward: TAAGAAGGAGATATACCATGAATGCTACGCCAACTAAGGGTGC
- Sirius Reverse: CTCGAGTGCGGCCGCAAGCTTAGCACCGGTGGAGTGACG
- BirA Forward: AAGGAGATATACATATGAAGGATAACACCGTGCCACTGA
- BirA Reverse: CTTTACCAGACTCGATTATTTTTCTGCACTACGCAGGGA
- BFP Forward: ACCACAGCCAGGATCCTATGAGCGAACTGATCAAAGAGAACA
- BFP Reverse: ATGCGGCCGCAAGCTTCTCATGCCATTCAATTTTCTGTGCT
- RFP Forward: AGGAGATATACCATGGCTTCCTCCGAAGACGTTATCAAAG
- RFP Reverse: GTGCGGCCGCAAGCTTAGCACCGGTGGAGTGACG
- Scygonadin Forward: GGCTGAAGCTGAATTCGGCCAGGCACTCAACAAAC
- Scygonadin Reverse: TGGGCCACGTGAATTCTCACTCATGCCATTCAATTTTCTG
Escherichia coli strains
Plasmid amplification strains- Stellar F-, endA1, supE44, thi-1, recA1, relA1, gyrA96, phoA, Φ80d lacZΔ M15, Δ(lacZYA-argF) U169, Δ(mrr-hsdRMS-mcrBC), ΔmcrA, λ-
- Top10 F- mcrA Δ(mrr-hsdRMS-mcrBC) φ80lacZΔM15 ΔlacX74 nupG recA1 araD139 Δ(ara-leu)7697 galE15 galK16 rpsL(StrR) endA1 λ-
- BL21(DE3) B F-ompT gal dcm lon hsdSB(rB-mB-) λ(DE3[lacI lacUV5-T7p07 ind1 sam7 nin5])[malB+]K-12(λS)
- Tuner F- ompT hsdSB (rB- mB-) gal dcm lacY1(DE3)
Pichia pastoris strains
For protein production we used GS200(ΔARG4, ΔHIS4) Strain for Cerberus production and X33(wt) for Scygonadin
Gluconacetobacter hansenii strain ATCC 53582
Click reagants
- DBCO-Biotin, Sigma CAS : 1255942-07-4
- DBCO-Fluorescein, Jena Bioscience Cat. No. : CLK-051-1
- DBCO-Magnetic beads, Jena Bioscience Cat. No. : CLK-1037-1
- Fluorescein-Azide, Jena Bioscience Cat. No. : CLK-80101-5
- 4-L-azidophenylanaline, Sigma Ref. : 06162
Graphene Functionalization
- Graphene, Sigma Ref. : 900561
- 4-Ethynylaniline, Sigma Ref. : 481122
- Isopentyl Nitrite, Sigma Ref. : 150495
- N,N-Dimethylformamide, Sigma Ref. : 227056
Protein purification
- Cobalt Affinity Gel, Sigma Ref. : H8162
- Protease inhibitor cocktail tablet, Sigma Ref. : S8830
Cellulose Binding assay
- Avicel, Sigma Ref. : 11365
No dogs were harmed over the course of this iGEM project.
The whole Toulouse INSA-UPS team wants to thank our sponsors, especially:









And many more. For futher information about our sponsors, please consult our Sponsors page.
The content provided on this website is the fruit of the work of the Toulouse INSA-UPS iGEM Team. As a deliverable for the iGEM Competition, it falls under the Creative Commons Attribution 4.0. Thus, all content on this wiki is available under the Creative Commons Attribution 4.0 license (or any later version). For futher information, please consult the official website of Creative Commons.
This website was designed with Bootstrap (4.1.3). Bootstrap is a front-end library of component for html, css and javascript. It relies on both Popper and jQuery. For further information, please consult the official website of Bootstrap.

