| Line 863: | Line 863: | ||
</p> | </p> | ||
<p> | <p> | ||
| + | <div class="image-container"> | ||
| + | <img src="https://static.igem.org/mediawiki/2018/d/d8/T--Vilnius-Lithuania--Fig1_NEW_thermoswitches.png"> | ||
| + | |||
<strong>Fig. 1</strong> Plots of translation efficiency vs. temperature. On the left hand side: plots of 10 modelled thermoswitches for Mstx-OmpA-GFP Nanobody. On the right hand side: plot of the selected thermoswitch to use with Mstx-OmpA-GFP Nanobody. RNA thermometer termed sw_6 displayed no artifacts, with near control-identical translation efficiency at high temperature and low efficiency at < 25 C. | <strong>Fig. 1</strong> Plots of translation efficiency vs. temperature. On the left hand side: plots of 10 modelled thermoswitches for Mstx-OmpA-GFP Nanobody. On the right hand side: plot of the selected thermoswitch to use with Mstx-OmpA-GFP Nanobody. RNA thermometer termed sw_6 displayed no artifacts, with near control-identical translation efficiency at high temperature and low efficiency at < 25 C. | ||
| − | </p> | + | </p></div> |
| − | + | <div class="image-container"><img src="https://static.igem.org/mediawiki/2018/c/ca/T--Vilnius-Lithuania--THERMO_fig_2.png"> | |
| + | <p> | ||
<strong>Fig. 2</strong> Plots of translation efficiency vs temperature. On the left hand side: plots of 10 modelled thermoswitches for GFP Nanobody-Iga-Mstx. On the right hand side: plot of the selected thermoswitch to use with GFP Nanobody-Iga-Mstx. RNA thermometer termed sw_5 displayed no artifacts, with relatively high translation efficiency at high temperature and largely lower efficiency at < 25 C. | <strong>Fig. 2</strong> Plots of translation efficiency vs temperature. On the left hand side: plots of 10 modelled thermoswitches for GFP Nanobody-Iga-Mstx. On the right hand side: plot of the selected thermoswitch to use with GFP Nanobody-Iga-Mstx. RNA thermometer termed sw_5 displayed no artifacts, with relatively high translation efficiency at high temperature and largely lower efficiency at < 25 C. | ||
| − | </p> | + | </p></div> |
| + | <div class="image-container"><img src="https://static.igem.org/mediawiki/2018/d/d8/T--Vilnius-Lithuania--THERMO_fig_3.png"> | ||
<p> | <p> | ||
<strong>Fig. 3</strong> Plots of translation efficiency vs. temperature. On the left hand side: plots of 10 modelled thermoswitches for Mstx-OmpA-His. On the right hand side: plot of the selected thermoswitch to use with Mstx-OmpA-His. RNA thermometer termed sw_8 displayed no artifacts, with near control-identical translation efficiency at high temperature and low efficiency at < 25 C. | <strong>Fig. 3</strong> Plots of translation efficiency vs. temperature. On the left hand side: plots of 10 modelled thermoswitches for Mstx-OmpA-His. On the right hand side: plot of the selected thermoswitch to use with Mstx-OmpA-His. RNA thermometer termed sw_8 displayed no artifacts, with near control-identical translation efficiency at high temperature and low efficiency at < 25 C. | ||
| − | </p> | + | </p></div> |
| + | <div class="image-container"><img src="https://static.igem.org/mediawiki/2018/a/a4/T--Vilnius-Lithuania--THERMO_fig_4.png"> | ||
<p> | <p> | ||
<strong>Fig. 4</strong> Plots of translation efficiency vs temperature. On the left hand side: plots of 10 modelled thermoswitches for His-Iga-Mstx. On the right hand side: plot of the selected thermoswitch to use with His-Iga-Mstx. RNA thermometer termed sw_4 displayed no artifacts, with relatively high translation efficiency at high temperature and largely lower efficiency at < 25 C. | <strong>Fig. 4</strong> Plots of translation efficiency vs temperature. On the left hand side: plots of 10 modelled thermoswitches for His-Iga-Mstx. On the right hand side: plot of the selected thermoswitch to use with His-Iga-Mstx. RNA thermometer termed sw_4 displayed no artifacts, with relatively high translation efficiency at high temperature and largely lower efficiency at < 25 C. | ||
| − | </p> | + | </p></div> |
<p> | <p> | ||
Thermoswitches were initially designed to appropriately melt and function at 37 C. Comparing the first curve in each plot which resembles the original sequence of our constructs without incorporated thermoswitch (control), it can be seen that novel designs show much stronger temperature dependence. However, they did not manage to achieve quite exact 37 C and displayed marginally lower translation efficiency than controls. Some sequences displayed artifacts that showed up as jumps in the efficiency plots. The believed reason was the usage of different SD sequences at low and high temperatures. For in vivo testing we selected designs that did not exhibit such jumps. Another interesting finding was that all thermoswitches designed for Iga protease bearing constructs showed a considerably lower efficiency of translation even at higher temperatures compared to OmpA bearing constructs, meaning that this characteristic was probably attributed to membrane protein structure and would be needed to be addressed in the future. | Thermoswitches were initially designed to appropriately melt and function at 37 C. Comparing the first curve in each plot which resembles the original sequence of our constructs without incorporated thermoswitch (control), it can be seen that novel designs show much stronger temperature dependence. However, they did not manage to achieve quite exact 37 C and displayed marginally lower translation efficiency than controls. Some sequences displayed artifacts that showed up as jumps in the efficiency plots. The believed reason was the usage of different SD sequences at low and high temperatures. For in vivo testing we selected designs that did not exhibit such jumps. Another interesting finding was that all thermoswitches designed for Iga protease bearing constructs showed a considerably lower efficiency of translation even at higher temperatures compared to OmpA bearing constructs, meaning that this characteristic was probably attributed to membrane protein structure and would be needed to be addressed in the future. | ||
| Line 881: | Line 887: | ||
The model was also applied to check the activity of thermoswitches that we have acquired from literature (see <a href="https://2018.igem.org/Team:Vilnius-Lithuania/Design">Design and Results</a>/<a href="https://2018.igem.org/Team:Vilnius-Lithuania/Design#RNA_Thermoswitches">RNA Thermoswitches</a>). Our model predicted fair, but viable switching effects for thermoswitch-GFP designs, which were later supported by in vivo measurements. | The model was also applied to check the activity of thermoswitches that we have acquired from literature (see <a href="https://2018.igem.org/Team:Vilnius-Lithuania/Design">Design and Results</a>/<a href="https://2018.igem.org/Team:Vilnius-Lithuania/Design#RNA_Thermoswitches">RNA Thermoswitches</a>). Our model predicted fair, but viable switching effects for thermoswitch-GFP designs, which were later supported by in vivo measurements. | ||
</p> | </p> | ||
| + | <div class="image-container"><img src="https://static.igem.org/mediawiki/2018/4/46/T--Vilnius-Lithuania--THERMO_fig_5.png"> | ||
<p> | <p> | ||
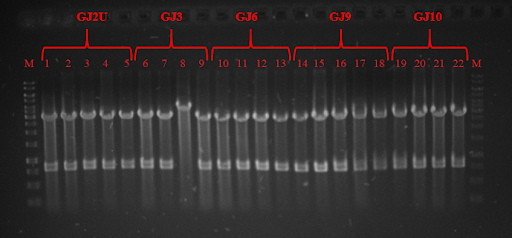
<strong>Fig. 5</strong> Plots of translation efficiency vs. temperature of the “GJ” thermoswithes-GFP constructs. Thermoswitches GJ2, GJ3, GJ9, GJ10 display similarly fair translation efficiency at 37 C, except for GJ6, which displays notably higher translation efficiency. GJ thermoswitches significantly differ in their activity at lower temperatures, with GJ9 locking the transcription most tightly and GJ3 being the leakiest of all tested designs. | <strong>Fig. 5</strong> Plots of translation efficiency vs. temperature of the “GJ” thermoswithes-GFP constructs. Thermoswitches GJ2, GJ3, GJ9, GJ10 display similarly fair translation efficiency at 37 C, except for GJ6, which displays notably higher translation efficiency. GJ thermoswitches significantly differ in their activity at lower temperatures, with GJ9 locking the transcription most tightly and GJ3 being the leakiest of all tested designs. | ||
| − | </p> | + | </p></div> |
<p></p> | <p></p> | ||
<h1>Model</h1> | <h1>Model</h1> | ||
| Line 890: | Line 897: | ||
A simple in-silico translation-initiation potential model<sup>5</sup> to quantify the likelihood of in vitro translation of a given mRNA sequence from a series of interaction energy parameters at constant temperatures was developed. The model defines the translation-initiation potential σ as: | A simple in-silico translation-initiation potential model<sup>5</sup> to quantify the likelihood of in vitro translation of a given mRNA sequence from a series of interaction energy parameters at constant temperatures was developed. The model defines the translation-initiation potential σ as: | ||
</p> | </p> | ||
| + | <div class="image-container"><img src="https://static.igem.org/mediawiki/2018/d/dd/T--Vilnius-Lithuania--THERMO_fig_6.png"> | ||
<p><strong>Fig. 6</strong></p> | <p><strong>Fig. 6</strong></p> | ||
<p> | <p> | ||
where R is the Boltzmann constant, T the temperature, ΔE<sub>SD</sub> the hybridization energy between the SD and anti-SD sequences, ΔE<sub>tRNA</sub> the hybridization energy of the start codon and its respective anti-codon (i.e, the tRNA<sup>Met</sup>), and ΔE<sub>open</sub> the energy required to unfold the 30-nucleotide-long RDS. Here, ΔE<sub>SD</sub> and ΔE<sub>tRNA</sub> are constant since neither the SD nor the start codon are altered. Consequently, variations in σ are exclusively determined by ΔE<sub>open</sub>. Applying the model to the plasmids with our constructs bearing thermoswitches, enabled us to rationalize translation events, as translatable constructs consistently scored higher σ, or lower ΔE<sub>open</sub>, than non-translatable ones. | where R is the Boltzmann constant, T the temperature, ΔE<sub>SD</sub> the hybridization energy between the SD and anti-SD sequences, ΔE<sub>tRNA</sub> the hybridization energy of the start codon and its respective anti-codon (i.e, the tRNA<sup>Met</sup>), and ΔE<sub>open</sub> the energy required to unfold the 30-nucleotide-long RDS. Here, ΔE<sub>SD</sub> and ΔE<sub>tRNA</sub> are constant since neither the SD nor the start codon are altered. Consequently, variations in σ are exclusively determined by ΔE<sub>open</sub>. Applying the model to the plasmids with our constructs bearing thermoswitches, enabled us to rationalize translation events, as translatable constructs consistently scored higher σ, or lower ΔE<sub>open</sub>, than non-translatable ones. | ||
| − | </p> | + | </p></div> |
<p></p> | <p></p> | ||
<h2>References</h2> | <h2>References</h2> | ||
Revision as of 19:19, 7 November 2018
Modeling
Mathematical model
Mathematical models and computer simulations provide a great way to describe the function and operation of BioBrick Parts and Devices. Synthetic Biology is an engineering discipline, and part of engineering is simulation and modeling to determine the behavior of your design before you build it. Designing and simulating can be iterated many times in a computer before moving to the lab. This award is for teams who build a model of their system and use it to inform system design or simulate expected behavior in conjunction with experiments in the wetlab











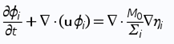


















 Fig. 1 Plots of translation efficiency vs. temperature. On the left hand side: plots of 10 modelled thermoswitches for Mstx-OmpA-GFP Nanobody. On the right hand side: plot of the selected thermoswitch to use with Mstx-OmpA-GFP Nanobody. RNA thermometer termed sw_6 displayed no artifacts, with near control-identical translation efficiency at high temperature and low efficiency at < 25 C.
Fig. 1 Plots of translation efficiency vs. temperature. On the left hand side: plots of 10 modelled thermoswitches for Mstx-OmpA-GFP Nanobody. On the right hand side: plot of the selected thermoswitch to use with Mstx-OmpA-GFP Nanobody. RNA thermometer termed sw_6 displayed no artifacts, with near control-identical translation efficiency at high temperature and low efficiency at < 25 C.