| Line 245: | Line 245: | ||
</div> | </div> | ||
</div> | </div> | ||
| − | </section | + | </section> |
Revision as of 11:22, 17 October 2018


From plastics to the power line
Polyethylene is the most widely used plastic and arguably one of the most versatile materials to ever be synthesized. Its practicality and convenience however, have come at a great environmental cost. Polyethylene takes millennia to decompose, leaching harmful microplastics into the environment. We approached this pressing issue from a synthetic biology perspective, making use of E. coli engineered with genes encoding for laccase to degrade polyethylene into smaller alkane chains. Our team recognizes the opportunity to further advance this project by addressing another key issue – energy. Using Shewanella oneidensis MR-1 strain’s inbuilt extracellular electron transport mechanism in tandem with genes responsible for alkane metabolism derived from Desulfatibacillum alkenivorans, we will generate electricity from the metabolism of degraded polyethylene, hoping that it will one day help in solving the world’s growing energy needs. Thus, our project serves as an integrated effort to simultaneously solve two crucial problems.
To achieve our goal of electricity generation from plastic degradation, we separated our project into three independent but interconnected modules. The first stage, the PE degradation module, focused on creating a genetic construct that would readily synthesize and secrete Laccase in order to degrade the polyethylene into smaller alkanes. Once fragmented, the alkanes would be processed by our second stage - the alkane metabolism module. This module plans to introduce alkane uptake and metabolism pathways into Shewanella oneidensis MR-1 so that it can process alkanes and channel the electrons obtained from its cellular respiration. The third and final module was the physical construct of the microbial fuel cell. We housed the Shewanella within a microbial fuel cell of our design to characterise and optimise the electricity generated. The following is a schematic to describe the flow of the project.


As a giant network of saturated hydrocarbons, polyethylene (PE) is a storehouse of a huge amount of chemical potential energy . While most microorganisms are unable to draw from this vast store as they cannot metabolise polyethylene or smaller alkanes, review of literature suggested that some microbes are capable of metabolising saturated hydrocarbons as their sole carbon source. There are however, limitations to the length of the carbon chains these microorganisms can metabolise [1].
With regards to our goal – to design a metabolism pathway for polyethylene, it was therefore a priority to fragment PE before it could be fed to the microbes. Further study on several research articles revealed that Rhodococcus ruber was among the few microorganisms that could utilize polyethylene as its main carbon source. Laccase was identified among several secreted metalloenzymes when the bacterium was grown on a PE film ([2], [6]) and so was suspected to be responsible for the cleavage effect on PE. Further analysis provided conclusive evidence that it was in fact Laccase cleaving the polyethylene backbone.
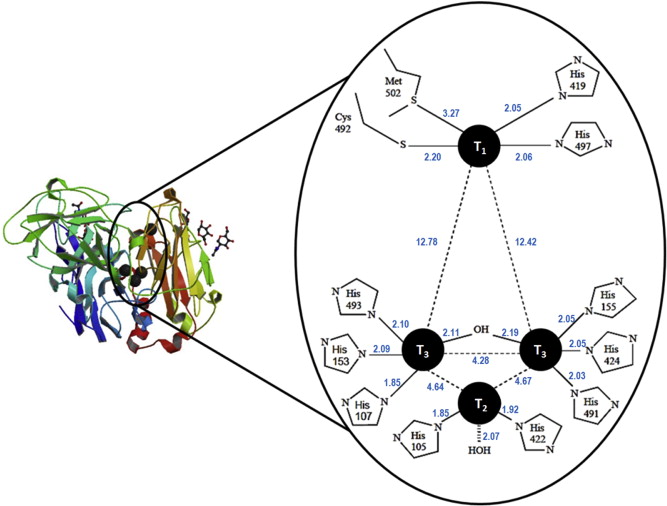
Figure 1. Protein structure and enzymatically active sites of laccase [7].
Laccase belongs to a family of enzymes known as multicopper oxidases which oxidize a variety of substrates while reducing dioxygen to water in the process. As shown in Figure 1, The enzymatically active part of laccase involves a cluster of 4 copper ions at different oxidation states [4]. The catalytic mechanism begins with the transfer of electron from substrate to the T1 copper site as a result of its higher redox potential; the electron obtained from the reduced T1 copper is then passed through the intermediate electron acceptor, T3 copper site and eventually ends up at the T2 copper site [4]. The T3 copper serves as an electron acceptor in the aerobic oxidation process while the presence of the T2 copper site is necessary as a terminal oxygen reducing site. Laccases were long reported to be present in some lignin- bio-degrading fungi, where they catalyse the oxidation, including carbonyl formation, of aromatic compounds. Nonetheless, there is considerable evidence showing laccase’s affinity to non-aromatic substrates, such as saturated hydrocarbons.
Due to the lack of any hydrolysable functional group, a specific enzymatic attack on the PE backbone may not be favourable; however, a random oxidation on PE could effectively weaken its chemical integrity. Presently, the activity of laccase on PE has been confirmed by documented cases from scientific research papers as well as from past iGEM projects. For instance, University London College IGEM team 2012 proposed that the laccase could lead to increased deterioration of polyethylene structure and was confirmed by the SEM (Scanning Electron Microscope) which showed an evident scratch made on a PE film surface after incubation with laccase, compared to the control. Moreover, in research conducted by other teams, crude laccase secreted by rhodococcus ruber was incubated with LDPE of average molecular weight 191,000. It reportedly resulted in 15 to 20 percent of mass reduction over a period of two weeks, which demonstrated the feasibility of laccase-degradation of polyethylene. To further enhance the efficiency of laccase, introduction of copper ions and mediators to the reaction medium were also justified to yield a higher degradation rate ([3], [5]) .
The target of this module was to demonstrate the ability of Laccase, expressed by E.coli, to fragment PE into simple hydrocarbons. For the purpose of utilizing laccase in the outer-membrane space, outer membrane protein A (OmpA) is required to serve as a protein signal to the membrane so that the laccase can be dumped to the intracellular space. Meanwhile, to minimise the misfolding of laccase with OmpA, our chosen OmpA includes a linker sequence at the N-termini to separate laccase from the signalling protein OmpA. To aid in protein extraction for characterization, a 6X his-tag region is added to all of our laccase. The final effective construct for activity assay is as follows:

Apart from the construct above, another construct was also involved to characterize the effectiveness of OmpA in bringing out laccase to the intracellular space.

With these two constructs constitutively expressed in E.coli, it is expected that we can compare the differences of laccase quantity in the culturing medium of E.coli for our characterisation.
References:
1. R. Wei and W. Zimmermann, “Microbial enzymes for the recycling of recalcitrant petroleum-based plastics: how far are we?,” Microbial Biotechnology, vol. 10, no. 6, pp. 1308–1322, 2017.
2. M. Santo, R. Weitsman, and A. Sivan, “The role of the copper-binding enzyme – laccase – in the biodegradation of polyethylene by the actinomycete Rhodococcus ruber,” International Biodeterioration & Biodegradation, vol. 84, pp. 204–210, 2013.
3. O. V. Morozova, G. P. Shumakovich, S. V. Shleev, and Y. I. Yaropolov, “Laccase-mediator systems and their applications: A review,” Applied Biochemistry and Microbiology, vol. 43, no. 5, pp. 523–535, 2007.
4. Jones, Stephen M., and Edward I. Solomon. “Electron Transfer and Reaction Mechanism of Laccases.” Cellular and molecular life sciences : CMLS 72.5 (2015): 869–883. PMC. Web. 13 Oct. 2018.
5. Fujisawa, M., Hirai, H., & Nishida, T. Degradation of Polyethylene and Nylon-66 by the Laccase-Mediator System. Journal of Polymers and the Environment, 9(3), 103-108, 2001
6. Hadar, & Sivan. Colonization, biofilm formation and biodegradation of polyethylene by a strain of Rhodococcus ruber. Applied Microbiology and Biotechnology, 65(1), 97-104, 2004
7. A. Piscitelli, C. Pezzella, V. Lettera, P. Giardina, V. Faraco, and G. Sannia, “Fungal Laccases,” Fungal Enzymes, Aug. 2013.

After the polyethylene plastic is decomposed by laccase, we plan to use bacteria to uptake the decomposed parts as a carbon source. As such, the second module of this project involves an alkane metabolism pathway, where the chassis organism is Shewanella Oniedensis MR-1, which can generate electricity for our project.
To generate the alkane metabolism pathway, the starting product (alkanes) must penetrate into the cytoplasm through the cell membrane. While short chain alkanes (less than C12) are able to diffuse directly into the cell, medium to long chain alkanes cannot due to their increased hydrophobic nature. Since the laccase fragmentizes PE into randomized lengths, it is in our interest to make sure that all ranges of alkane size can be efficiently metabolized within the cytoplasm. To this end, we have to use an outer membrane alkane channel protein to mediate the entry of the medium to long chains alkane.
According to the literature, an outer membrane channel protein called AlkL from Pseudomonas putida GPo1 has been shown to enhance the diffusion efficiency of dodecane (C12) into the cell when expressed in Escherichia coli. It is also shown to be able to facilitate the transport of larger alkanes, which has high hydrophobicity, making it more difficult to pass through the lipopolysaccharide outer membrane of the bacteria. The mechanism behinds AlkL is that it penetrates through the lipopolysaccharide layer, where an extracellular domain with high affinity for hydrophobic molecules lies. After the alkane molecule is collected via this domain, it makes its way through the hydrophobic core of AlkL . This subsequently allows the alkane to makes its way to the cytoplasm via a small lateral opening of AlkL. From there, the alkane may make its way to the cytoplasm. The outer membrane alkane channel protein(AlkL) we plan to use is derived comes from Pseudomonas Oleovarans. However, there is one drawback with AlkL: when overexpressed, it is toxic to the host cell. As such, we will attempt to express it with different strengths of promoters and test the effect of AlkL toxicity on the host’s growth rate before proceeding.
After successfully passing through the bacterial cell wall barrier, alkane can undergo degradation. Originally, it was thought that alkane degradation could only occur in the presence of oxygen. This was soon proven to be false: discovered in a bacterial species known as Desulfatibacillium alkenivorans AK-01 found originally in water contaminated with oil spills, its native alkane degradation can operate in anaerobic conditions. It is also able to metabolize the medium to long chain alkanes. This feature is key to developing the further step of electrogenicity in the MFC part of our project.
Desulfatibacillium alkenivorans AK-01 had its genome sequenced in 2012 and a theoretical model for its alkane degradation pathway involving 4 major enzymes was proposed, which we hope to use in the second part of our project. The pathway is as follows: once the alkane fragment enters the cell cytoplasm, it combines with fumarate under the action of alkylsuccinate synthase (ASS) to form 1-methylalkylsuccinate. Then, CoA Transferase adds a CoA group to it, producing 1-methylalkylsuccinyl-CoA. Afterwards, its C-skeleton is rearranged by methylmalonyl-CoA mutase to be converted it to 2-methylalkylmalonyl-CoA. Finally, it is converted to 4-Methyloctadecanoyl-CoA by Carboxyl transferase where a CO2 is removed. Upon reaching this stage of molecular rearrangement, the compound is ready to enter β-oxidation of the cell.

Our design incorporates the 4 above steps into Shewanella which is hypothesized to functionally metabolize alkanes. It is hoped that once the alkane can ultimately enter β-oxidation, it will subsequently enter the TCA cycle by the cell’s natural processes so that it can serve as a carbon source for Shewanella to generate electricity. However, some of the aforementioned proteins are actually made from subunit peptides. Therefore, to enable such a metabolic pathway, theoretically a total of 9 genes are necessary. The genes AssA1, AssB1, AssC1, AssD1 code for subunits that form alkylsuccinate synthase, assisted by AssE1 that encodes for a chaperone-like protein. AssK1 encodes for CoA transferase. McmSS and McmLS code for methylmalonyl-CoA mutase. And finally, Dalk_1740 codes for Carboxyl Transferase. The original plan was to transform all of them into Shewanella but the complexity of the genes posed a major challenge, one which we intend to solve next year.
Therefore, by engineering AlkL and the alkane metabolic pathway into Shewanella, we hope it provides a bridge between PE degradation and exo-electrogenicity.
References:
1. Grant, C., Deszcz, D., Wei, Y., Martínez-Torres, R., Morris, P., Folliard, T., Sreenivasan, R., Ward, J., Dalby, P., Woodley, J. and Baganz, F. (2014). Identification and use of an alkane transporter plug-in for applications in biocatalysis and whole-cell biosensing of alkanes.
2. Callaghan, A., Morris, B., Pereira, I., McInerney, M., Austin, R., Groves, J., Kukor, J., Suflita, J., Young, L., Zylstra, G. and Wawrik, B. (2011). The genome sequence of Desulfatibacillum alkenivorans AK-01: a blueprint for anaerobic alkane oxidation.
3. Herath, A., Wawrik, B., Qin, Y., Zhou, J. and Callaghan, A. (2016). Transcriptional response of Desulfatibacillum alkenivoransAK-01 to growth on alkanes: insights from RT-qPCR and microarray analyses.

