Golden Gate Assembly is a revolutionary cloning method published 10 years ago (Engler et al., 2008). since its publication it became very popular giving rise to many assembly strategies based on this system. All systems published over the years offered improvements over previous publications. Nevertheless, although there are many systems available – often specialized for certain organisms or classes – the development by far is not at an end. We worked on a novel toolbox and developed an improved, highly flexible system focused on prokaryotes including a high number of parts characterized for the novel upcoming chassis V. natriegens.
The innovation given by Golden Gate assembly was the introduction of type IIS restriction enzymes used for DNA digestion. A current workflow at that time consisted of DNA restriction, gel purification and finally ligation of the single parts. For restriction different type II restriction enzymes were taken making it important to check for compatible buffers and getting a six base pair scar after ligation. By introducing type IIS restriction enzymes into this workflow the system was highly improved. This new method offered many advantages like I) an independent enzyme recognition site that is removed after restriction, II) restriction and ligation in one pot III) directional assembly by conscious design of the cleavage sequences IV) no buffer incompatibilities due to the usage of one enzyme cutting at several spots and still producing different cleavage sequences (Engler et al., 2008) (Figure will be added…).
Based on this, many new systems arose offering the possibility of building complex constructs out of single genetic parts in a fast, simple and flexible way. Full libraries of standardized genetic parts were developed enabling the user to build full combinatorial libraries. The first system developed was the modular cloning system (MoClo) offering a new hierarchical cloning method of defined biological parts (Weber et al., 2011) and consequently setting a new standard over the years.
Here we summarize some Golden Gate based systems and compare their advantages and disadvantages.
Image description
The modular cloning system was the first one proposing a standard for Golden Gate based assembly. This toolbox offers 5 types of modules designed mainly for eukaryotes. The modules are stored in level 0 acceptor plasmids derived from the pUC19 backbone with a spectinomycin resistance and a LacZα cassette as dropout for blue/white screening. Custom level 0 plasmids are assembled by flanking the sequences with BpiI recognition sites and setting a single restriction-ligation-reaction with the correspondent plasmid. Up to 5 level 0 modules are assembled in an acceptor plasmid with ampicillin resistance and a LacZα-dropout by restriction with BsaI to transcription units. 6 transcription units can be assembled using BpiI into level 2 multigene constructs containing a kanamycin resistance and a Cred-dropout. Alternatively Esp3I can be use to transfer the constructs to intermediary levels to reach higher levels for assembly of bigger constructs.
The first CRISPR-loci were discovered in 1987 in the E. coli strain K12 (Ishino et al. 1987) and the first description of a cas gene and their homologies to DNA-related enzymes appeared in 2002 (Jansen et al. 2002). Since then, more than 9000 research articles have been published about the different CRISPR/Cas subtypes and their application in various CRISPR based technologies (Adli 2018). The potential applications are almost limitless, ranging from new methods in research to industrial applications as well as human gene therapy. With CRISPR/Cas, it is possible to specifically switch off genes in a large number of prokaryotic and eukaryotic organisms, to integrate foreign DNA into the genome of an organism or to use the DNA-binding property for in vitro experiments (Hsu et al. 2014).
Due to its simplicity and modularity, the Type II CRISPR/Cas9 system from Streptococcus pyogenes is a commonly used tool for genetic engineering applications in a variety of organism (Hsu et al. 2014, Charpentier & Marraffini 2014, Doudna & Charpentier 2014). For its RNA-guided silencing of foreign DNA it requires just one Cas9 encoding gene and the action of the CRISPR RNA (crRNA) as well as the trans-activating crRNA (tracrRNA) (Jinek et al. 2012, Gasiunas et al. 2012). During infection, the CRISPR-locus will be transcribed into the pre-crRNA containing the spacer region flanked by conservated repeats. The tracrRNA binds to the pre-crRNA due to 25 nucleotides which are complementary to the repeat regions of the pre-crRNA (Deltcheva et al. 2011). After procession, the mature crRNA:tracrRNA duplex binds the Cas9 with the tracrRNA builded stem-loop constituting a CRISPR ribonucleoprotein (crRNP). The crRNA contained a 20 nucleotides large spacer region at the end of the N-terminus which is complementary to the invading DNA (protospacer) therefore leading the crRNP to the target. In addition, the presence of the Protospacer adjacent Motif (PAM, NGG in S. pyogenes) preceding the protospacer three nucleotides is necessary for the cleavage activity (Jinek et al. 2014, Nishimasu et al. 2014). The crRNA and tracrRNA can directly be linked by a linker loop into a single guided RNA (sgRNA) like we used in our project for a more simplified and modularized genetic engineering application (Brazelton et al. 2016, Jinek et al. 2012).
Our Goal
Our goal is to accelerate metabolic engineering by establishing a workflow for fast pathway construction, product screening and pathway optimization. Metabolic engineering is a broad field in synthetic biology, where existing pathways are divided into modular parts and then used and combined to build new pathways ( (Stephanopoulos et al.2012) ). One of the best examples for the impact of metabolic engineering on solving global problems is the engineered production pathway for artemisinic acid, the precursor for a drug against malaria (Ro et al.2006) . In the old days the drug had to be isolated out of the plant Artemisia annua. This was very inefficient because of the limitations of plant growth and low yields of artemisinic acid. Therefore, the group of Jay Keasling engineered a metabolic pathway to produce the drug in yeast cells, which grow faster and produce a lot more artemisinic acid. As in this example, metabolic engineering enables the production of drugs and other valuable products with an increased yield when compared to product isolation from natural sources. Furthermore, the large variety of usable enzymes, pathways and chassis enables more flexibility in pathway design compared to chemical approaches.
Why should we accelerate metabolic engineering?
Although much progress in metabolic engineering was made in the last 20 years, there are still many drawbacks and limitations.
For instance, it is crucial to have a big set of genes as combinable parts to find the optimal pathway and they have to be predictable.
With many of them just poorly characterized, rational pathway design remains difficult.
Our understanding of the working and interactions of enzymes at the level of the metabolism is still rudimentary. Consequently, metabolic engineering consists of much trial-and-error and researchers have to endure series of failures.
One strategy to face these issues is modeling of enzyme behavior but currently limitations in computing power prevent this from being used in high-throughput. Another strategy is to use chassis as workhorses to test a lot of enzyme combinations and pathway variants. Here, organisms like E. coli, S. cerevisiae or C. glutamicum are often used but product formation is limited by growth rate and nutrient uptake
(Stephanopoulos et al.2007).
When a pathway is constructed and to be tested, one has to wait until the production strain has grown and then product formation is still limited by substrate uptake. Another big drawback is that screening for product often is expensive, time consuming and complicated.
Conventional methods like LC-MS need a lot of time and resources and it is hardly possible to use it in high-throughput, for instance for a whole pathway library.
To overcome these limitations and accelerate metabolic engineering, we established V. natriegens as chassis for metabolic engineering.
Its doubling time of under 10 minutes makes it the perfect organism to quickly test many pathway variants or enzyme versions.
Additionally, it has a glucose uptake rate of 3.90 g g-1h-1 while E. coli, S. cerevisiae and C. glutamicum have much lower rates (1.90, 3.52 and 0.37 g g-1h-1, respectively).
Coupled with the short doubling time, V. natriegens is a powerful chassis not only for research purposes but also for industrial usage, where high productivity is crucial.

How are we accelerating metabolic engineering?
For optimal execution of metabolic engineering we established a workflow, basing on the DBTL-cycle. We use our Marburg Toolbox to build whole pathway libraries, which can then be transformed into our producer strain VibriXpress. By using biosensors for product screening, we circumvent the expensive and time-intensive usage of LC-MS and learn which pathway variants are the most promising. These findings can then be fed into our pathway-design, thereby closing the DBTL-cycle. To further optimize these pathways, we planned to use directed evolution for more adaptations to that route.
3-Hydroxypropionic acid 3HPA
As a proof of concept for our engineering workflow we chose the biological production of 3-Hydroxypropionic acid ( 3HPA ). 3HPA is a compound of high industrial value.
In 2004 the U.S. Department of Energy recommended 3HPA as alternative to fossil oils for the chemical industry
(Werpy et al.2004).
3HPA as a platform chemical can be converted into many other compounds like acrylic acid and acrylamide, which both are also precursors for further compounds.
According to an estimate of 2014, acrylates alone have an annual market value of USD12 billion, making its precursor 3HPA an optimal target product for our metabolic engineering approach.
By finding the optimal pathway to produce 3HPA we could show the efficiency of metabolic engineering in V. natriegens as well as getting a big step closer to an alternative to fossil oils by biological, renewable resources.

Our pathway
In recent years many people concentrated on producing 3HPA via many different metabolic pathways
(Valdehuesa et al.2013)
;
(Vidra et al.2017).
Most of these pathways use glycerol or glucose as starting substrate, but there are also publications where acrylic acid, CO2 or uracil were converted to 3HPA
(Vidra et al.2017).
One promising route starts with glycerol which is dehydrated and oxidized.
For E. coli, a productivity of 6.6 mmol g-1 cdw h-1 (oder Lieber andere units? 6.5mmol l-1 oder 0.48 mol mol-1 glycerol) has been shown
(Raj et al.2008).
However, this route depends on vitamin B12, which often can’t be taken up or produced by the chassis.
Many other routes start with glucose, which gets metabolized via propionate, lactate, β-alanine or malonyl-CoA
(Vidra et al.2017).
We decided to use a pathway, based on the conversion of acetyl-CoA into malonyl-CoA and finally 3HPA . According to
(Valdehuesa et al.2013)
who evaluated many pyruvate-derived production pathways from a thermodynamic point of view, this is one of the most efficient routes for 3HPA production.
This pathway was first established in E. coli, but it was also tried in many other chassis like S. cerevisiae and S. elongatus
(Kildegaard et al.2016);
(Liu et al.2017);
(Rathnasingh et al.2012).
We chose this route for several reasons.
Firstly, it is based on glucose degradation and V. natriegens has an unbeaten glucose-uptake rate, enabling high glucose consumption. Secondly, vitamin B12 is not necessary, the only cofactors involved are NADPH and biotin, both of which occur naturally in the organism.
Thirdly, theoretically just one further enzyme is needed to complete the pathway. All enzymes necessary for the production of malonyl-CoA are already present in V. natriegens and we just have to integrate the last enzyme, malonyl-CoA reductase.
Nevertheless, overexpression of acetyl-CoA carboxylase increases production, so we took both enzymes into consideration.
(Liu et al.2017)
proposed, that there are many ways to further optimize the pathway and direct the flow towards a high product titer.
Hence, implementing many variations of this pathway into V. natriegens and testing them in a fast manner promises a wealth of knowledge about pathway optimization and general principles of metabolic engineering.


One major goal of synthetic biology is to make biology engineerable and predictable. Currently metabolic engineering does still involve a lot of trial and error. With the number of possibilities for pathway finetuning this scales very fast, becoming very difficult to tackle with traditional methods.
Also, once we create such a library of metabolic pathway variants, possibly including thousands of strains, we have to find the most promising ones among these. A traditional approach would be to check the production of each strain in its own culture flask. Using for example HPLC or mass spec.
A more suitable, easy way to quickly screen for metabolite concentration in a semi quantitative way are biosensors. One of the easiest to use classes of biosensors are metabolite-responsive transcription factors
(Liu et al. 2018).
They are usually very specific towards their distinct substrates. In vivo, most of these systems help their native organism to react to environmental clues by regulating expression of for instance catabolic enzymes or importers.
In the past, these have ben co-opted by researchers to function as sensors or to control enzyme expression to regulate a pathway depending on the availability of a certain metabolite
(Rogers, Taylor, and Church 2016).
By using reporter genes like GFP or the LUX operon, we have a measurable output that can be used to select for the best contenders. By running this in several iterations, nature's most powerful optimization tool, evolution, can be applied to further improve the pathway
(Williams et al. 2016).
Applying fluorescence activated cell sorting (FACS) coupled with
V. natriegens
incredible doubling time, thousands of generations can be screened for progressively better pathway variants.
So, we decided to put our toolbox full of characterized parts to good use. The Marburg collection contains all the parts needed to find the ideal enzyme balance for highest production. Being able to build a large number of differentially assembled pathways is the answer to creating the most productive strain. Finding the right one in that complexity is another difficulty to overcome, one we solved by finding transcriptional regulators that specifically activate reporter gene expression relative to the concentration of key metabolites, 3-hydroxypropionate and Malonyl-CoA. This gives us a quantifiable output with which we are able to judge the different combinations. Also, utilizing the speed of V. natriegens , the time required for a high throughput screening can be reduced dramatically.
We deemed the concentration of two different metabolites as being critical to our project: 3-Hydroxypropionate, the product itself, and malonyl-CoA, an important intermediate we also plan to overexpress.

Using the information from the paper of
Hanko et al. (2017),
we settled for the
hdpR
/
hdpH
system coming from
Pseudomonas putida
KT2440. It consists the constitutively expressed HdpR protein
(BBa_K2560304).
, which is able to bind to the P
HdpR
and induce expression of a gene. In the native setting, that would be the
hdpH
gene, encoding for a 3-hydroxypropionate dehydrogenase which enables
P. putida
to grow on 3HPA . We cloned our reporter gene in its position. This way we got a direct readout of the 3HPA concentrations in the individual cells.
Because our model showed that the low activity of
V. natriegens
native Acc , malonyl-CoA is likely to be a limiting factor in 3HPA production. Having a direct indication to how much malonyl-CoA is available

Directed evolution
Although the Marburg Toolbox enables the construction of many pathway variations, there are still many more possibilities for direct metabolic fluxes towards 3HPA . To enforce these flux improvements, we planned to use directed evolution coupled with FACS selection. By implementing genes for the conversion of 3HPA into succinate, we lead our product back to the central metabolism. By deleting all other genes responsible for succinate production, the cells are forced to use the 3HPA bypass to get succinate, which is needed in the TCA cycle. The big advantage of using V. natriegens for that purpose is that it’s short doubling time increases the number of cell divisions per time and thereby increases the number of mutations in a given time frame. Coupled with our biosensors and fluorescence-activated cell sorting (FACS) we can easily and rapidly select for cells performing the most efficient pathways. Additionally to mutation-guided evolution there are many genome modifications which could increase flow towards 3HPA , for instance by downregulation of side reactions. In the Design-Section a list of possible genome modifications is shown. [Link zu Directed evolution im Designabschnitt machen]. By using MAGE , it is possible to get a library of different genome modifications, which can then be selected via FACS for their efficiency to produce 3HPA or succinate. MAGE stands for multiplex automated genome engineering and is a technique to create combinatorial genome modifications. MAGE uses a set of oligos, which get transformed into a population of cells and then integrate into the chromosome via homologous recombination. Different cells in the population will have different numbers and combinations of modifications leading to a genomic diversity. By applying MAGE in combination with FACS for our pathway we get a huge variety of possible pathway optimizations which can be sorted via FACS based on their efficiency.
Applications of our workflow
By using V. natriegens as a chassis for metabolic engineering, it is possible to build, test and optimize pathways and enzymes at a high-throughput level. The Marburg toolbox enables the fast construction of pathway libraries, while the genetic accessibility of the strain makes genome modification and pathway integration simple. The usage of our biosensors for screening of malonyl-CoA and 3HPA accelerates identification of the best pathway variants and the short doubling time coupled with the high nutrient uptake rate empowers V. natriegens to be the perfect chassis for metabolic and enzyme engineering!
First isolated from salt marsh mud on Sapelo island off the coast of Georgia in 1958 (Payne et al. 1958), the gram-negative bacterium was first named Pseudomonas natriegens (Payne et al. 1960). These early studies also revealed a broad range of tolerated pH conditions with an optimum at a pH of 7.5 (Payne et al. 1960). One feature of this newly discovered organism became immediately apparent: The incredible doubling rate, first determined to be around 9.8 minutes (Eagon 1962). We found that doubling rates of around 7 minutes were possible (own Data).
Only much later, studies found several possible reasons for this. Firstly, as most Vibrio species, V. natriegens genome is split and distributed on two chromosomes. This means that replication can start at two origins of replication (Ori), resulting in more in parallel replication.
Additionally, it was shown that the gamma-proteobacterium has an increased rRNA activity. This is because of a greater number of rRNA operons which are additionally controlled by stronger promoters (Aiyar et al. 2002) when compared to E. coli . More rRNA means more ribosomes since rRNA synthesis has been shown to be the rate-limiting step in their assembly (Miura et al. 1981). Estimates for the number of ribosomes in V. natriegens in the exponential phase suggest around 115,000 per cell, while E. coli is estimated to have between 70,000 and 90,000 (Failmezger et al. 2018).
This gives V. natriegens higher protein expression and lets it create biomass more quickly. Could this also be exploited for the production of high-value proteins? Most likely.
It was soon discovered that V. natriegens would make a perfect example by which to effectively and harmlessly demonstrate the basic techniques of microbiology to students, taking full advantage of the immense doubling rate (Mullenger 1973; Delpech 2001). Experiments regarding population structure, UV stress, and simply bare growth can be observed in a much shorter time frame.

The rod-shaped bacterium occupies a special ecological nice in estuarine and salt marsh regions, where strongly fluctuating salt concentrations and nutrient availability create a challenging environment. In order to adapt to these challenges, V. natriegens can utilize a wide range of substrates to outgrow competitors, as well as remain in a low metabolic state for extended periods of time when the nutrient pool is depleted (Nazly 1980). Furthermore, it has the ability to quickly draw available phosphorous from its environs and store them in polyphosphate bodies (Nissen et al. 1987) for later utilization. Yet another way in which V. natriegens is adapted to its variable environment is the storage of carbon as polyhydroxyalkanoates (PHAs), biodegradable polyesters that are accumulated in intracellular granules that can be broken down under starvation conditions (Chien et al. 2007).
From the well oxygenated upper layer of mud down to the anoxic layers, V. natriegens is able to grow in many surroundings. These extremes are challenges to which V. natriegens found solutions. At the surface of the mud, as well as in open waters, the bacteria are exposed to strong UV radiation. As a response to this, the DNA damage repair systems are highly active (Simons et al. 2010), with significantly elevated expression levels of some parts of the system compared to E. coli , thus increasing DNA sequence integrity. By switching to fermentation, growth can be maintained under anaerobic conditions, allowing the colonization of the lower strata in the mud.
V. natriegens was shown to be a vital part of the marine ecosystem because it has the rare ability to fix atmospheric nitrogen under anaerobic conditions, thereby enriching its habitat and playing a crucial role as a provider of essential, bioavailable nitrogen (Coyer et al. 1996) under anaerobic conditions.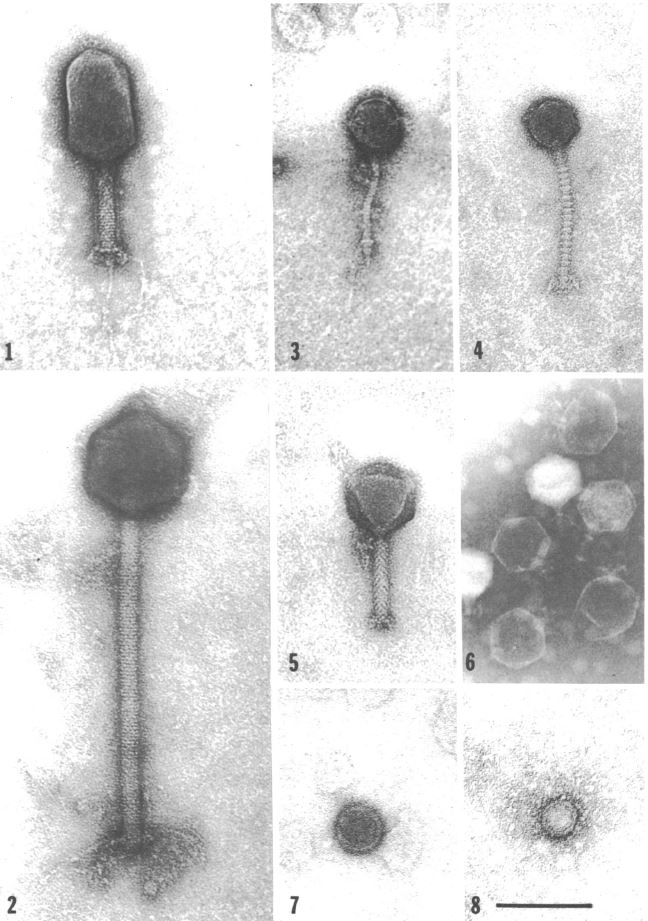
Unsurprisingly, V. natriegens has its own predators in its native habitat: Bacteriophages (Zachary 1974). Studies in the 70ies found a link between different phages and the prevailing environmental conditions (Zachary 1978) . This suggests the phages employ a divide and conquer strategy, specializing, thereby limiting competition while improving reproductive success. One phage may have adapted to be effective at lower temperatures, while others prefer higher temperatures. The same applies to other conditions like for instance salt concentration (Zachary 1976). Intriguingly, that opens the possibility of creating genetic parts derived from these phages, applicable to future work with V. natriegens, analogous to the T7 system in E. coli. On the other hand, one study could show, that the phage carrying the toxins associated with V. cholerae could not replicate in V. natriegens (Lee et al. 2016), further showing its harmlessness towards humans.
In contrast to that, some strains of Vibrio natriegens seem to be predators themselves. Confirmed as being pathogenic in several marine crustacea, most notably in the swimming crab Portunus trituberculatus, in which V. natriegens can lead to mortalities up to 85% (Bi et al. 2016). This is especially problematic since P. trituberculatus is farmed commercially in aquaculture in south-east China. The mechanism of that infection is poorly studied and it is not clear whether all strains of V. natriegens are pathogenic to crustaceans, but it is well known that many Vibrio species cause opportunistic infections in crustacea.
On a more positive note, a recent study could show that the sponge Aplysilla rosea develops a more diverse microbiome when exposed to V. natriegens (Mehbub et al. 2018). It can be speculated that this helps to protect the sponge from pathogenic microbes that then find it harder to colonize its surface.
A diverse, well-balanced metabolism allows a bacterium to grow in many conditions, while simultaneously making it interesting for biotechnological applications. V. natriegens can utilize a wide range of substrates (Hoffart et al. 2017), as well as grow in the absence of oxygen while being faster than other bacteria under the same conditions.
For many applications in which a modern laboratory work horse is expected to be useful, a high-resolution map of its metabolism is a prerequisite. One recent study generated just that (Long et al. 2017). By tracking 13C labeled carbon through the intricate network that constitutes V. natriegens metabolism, they were able to gain insight. Fore one, they found the it to be very similar to the well-studied E. coli metabolism. It has at the major catabolic and anabolic pathways, the same canonical amino acids and building blocks. Some notable differences between the two were the RNA content being 29% in V. natriegens compared to only 21% in E. coli, the presence of an enzyme for the decarboxylation of oxalate (Long et al 2017).
The specific uptake of many carbon sources per gram dry weight per time was sown to be significantly higher than in other comparable species of bacteria (Hoffart et al. 2017). Many commercially interesting compounds are already produced by unmodified V. natriegens. In the case of alanine production, only four deletions resulted in a strain that outperformed highly specialized E. coli and C. glutamicum production-strains by a factor of 9 to 13 times (Hoffart et al. 2017). Bioreactor based production on a large scale is expensive, therefore a strain generating similar or even higher yield per gram carbon source is highly desirable, improving temporal yield per fermentation unit.
When growing anaerobically, acids are produced that lead to an acidification of the medium but when grown under strong aeration, the pH can, depending on the medium, rise. This is most likely due to formation of ammonia (Eagon 1962).
The dependence on Na+ ions for its metabolic activity conveys a form of natural biocontainment, inhibiting growth in accidentally released V. natriegens (Webb and Payne 1971). The high salt content of the LBv2 media also presents possible contaminants with a barrier, reducing the likelihood of airborne microbes to take hold.
Recently, a study revealed that V. natriegens, surprisingly, is the most effective producer of selenium nanoparticles yet described (Fern`ndez-Llamosas et al. 2017). These particles with a diameter from 100-400nm have applications in diverse fields, such as medicine, microelectronics and more. Producing the nanoparticles in living systems has benefits over other methods, for instance, the low energy consumption as well as a coat of proteins preventing agglomeration of the particles. Also, V. natriegens exceptionally high resistance to selenite suggests possible applications in bioremediation of contaminated water and soils.
Working with V. natriegens
ATCC 14048 is the most commonly used strain of V. natriegens. Most results from earlier studies were generated from this strain. We recommend using ATCC 14048 in order to ensure that the findings will be applicable to your work and to adopt it as the standard strain. It is also the fastest strain known (Weinstock et al. 2016) so you won´t miss out on its incredible doubling time.
Stemming from subtropical regions, V. natriegens has not the same tolerance to low temperatures that we are used to from E. coli, but it is still much more resilient to cold than B. subtilis. At the root of that lies the low catalytic activity of V. natriegens native catalase at low temperatures, which is then unable to detoxify reactive oxygen species. To address that problem, scientists from SGI introduced homologous catalases, thereby creating a strain with comparable cold tolerance. With this adaptation, storage on plates can now be extended beyond the four weeks at room temperature by placing the plates in the fridge (Weinstock et al. 2016) In storage at -80°C in a standard 20% glycerol stock the cells stay viable almost indefinitely. And even when working with the wildtype, your cells won´t die overnight when placed at 4°C. Only after a week they will notably start decreasing in viability.
When working with V. natriegens ATCC 14048, no special precautions have to be taken since it has been shown that no known pathogenicity associated genes were present (Weinstock et al. 2016) Also, there is continued record of research since 1958 with not a single documented incident of a human infection with V. natriegens.
Lately, a new branch of microbiologists developed an interest in V. natriegens. Their main focus lies in establishing state of the art methods and bringing V. natriegens to its full potential. We see our project as part of this movement and wish to make V. natriegens easily accessible to researchers. One groundbreaking paper (Weinstock et al. 2016) established several methods to make V. natriegens genetically accessible, as well as characterizing some central genetic parts. This paved the way for more research into applying its potential.
If you want to introduce genetic material into V. natriegens, you have several tried and tested options to choose from. You could use electroporation, a quick and simple method and the one with the highest transformation efficiency. Electroporation can also be used on wildtype cells. Or you could use chemically competent cells, the transformation works very similar to E. coli heat shock transformation. But in order to achieve good efficiencies using chemically competent cells, a nuclease deletion strain is needed. And if you want to insert especially large constructs, you can also use conjugation with very good success ( (Lee et al. 2016; Weinstock et al. 2016).
A fully sequenced genome is also a prerequisite for many studies. Currently, several sequences are available (Lee et al. 2016; Maida et al. 2013), one of them generated by us. The genome has a size of approximately 5.17 Mb. Automated annotation was performed, revealing more of the genomic structure, as well as enabling the reconstruction of the codon usage profile (Lee et al. 2016).
The genome of V. natriegens is, like in many species of the genus, split into two chromosomes. Chromosome one contains 3.24 Mb, chromosome two 1.92 Mb.
Plasmids, Promoter, resistance
Usually, a big hurdle when changing your chassis organism is a very basic question: Will my constructs still work? Do I need different ori? Do I have to redesign everything from scratch? Fortunately, the most commonly used origins of replication, ColE1, p15A and pMB1 are maintained just fine in V. natriegens (Weinstock et al. 2016) We could observe some differences in copy number, which we quantified. You can find the corresponding data on our Part collection page. To retrieve these plasmids from the cells, we have tested a wide variety of commercially available kits and found that all of them worked, returning plasmid of the same purity and yield as for E. coli. Promoter commonly used in E. coli also lead to expression in V. natriegens. Expression levels are slightly different, but we also quantified these and you can find the results on our toolbox page. The same goes for other basic genomic parts like terminators and ribosomal binding sites.
Commonly, when very high gene expression is needed, for instance in protein overproduction, the phage derived T7 system is used. The T7 phage infects E. coli but it was also successfully inserted into V. natriegens and shown to drive protein expression (Weinstock et al. 2016) One way of generating even stronger, phage derived promoters could be using V. natriegens derived phage expression systems.
Every organism has its unique set of resistances to certain antimicrobial molecules. In order to successfully use a new organism, awareness of its individual tolerances is required to use antibiotics successfully for selection. We could show that chloramphenicol, carbenicillin, tetracycline and kanamycin worked, but at different concentrations from E. coli. We measured the V. natriegens specific concentrations and made them available on our strain engineering page.
Plasmids in absence of a selective pressure were only retained for a short time (Own Data). Curing of plasmids that have no longer a purpose is therefore quick and fairly effortless.
MuGENT - Recombineering
Investigations into alternative genetic tools were also undertaken, most notably MuGENT and Recombineering.
MuGENT takes advantage of V. natriegens natural competence by inducing the TfoX system. The TfoX system, which is known from many Vibrio species, enables uptake and genomic integration of DNA parts via homologous recombination (Pollack-Berti 2010). In most cases, growth on chitin was identified as the trigger which activated the system. Regrettably, the trigger is not known for V. natriegens. Using this system, a group of researchers characterized this system in V. natriegens, making it usable by placing it under the control of an inducible promoter. Additionally, they demonstrated its potential by increasing the natural poly-hydroxybutyrate (PHB) production 100-fold (Dalia et al. 2017).
Recombineering takes advantage of the fact that short DNA oligonucleotides, that can easily be delivered into cells by a plethora of ways, can be used to generate point mutations, knockouts and small modifications in the genome, as well as on plasmids. The homologous flanks required are relatively short, allowing for constructs under 100 bp. Having this system available in V. natriegens means that generating new strains and gene variants is easy and quickly implemented (Lee et al. 2017).
Cell Free Protein-expression Systems (CFPS)
Very recently, several groups have published their work about cell free protein expression systems (CFPS) in V. natriegens (Failmezger et al. 2018; Des Soye et al. 2018; Wiegand et al. 2018). Cell free systems offer the advantage of direct access to many cell components as well as fine control over the conditions in the reaction (Carlson et al. 2012). While these are true for all CFPSs, the advantage of creating such a system from V. natriegens is the much higher ribosome count (Failmezger et al. 2018).
Although the system works well when expressing plasmids from E. coli, it could be shown that expression was much higher when plasmids prepared from V. natriegens were used. Possible explanations are differences in methylation patterns, resulting in degradation of the plasmid by the system (Failmezger et al. 2018). All this could be done by using protocols very similar to well established methods for other bacteria.
Cre- lox
The Cre- lox system was also successfully implemented and shown to work consistently with 600bp homology flanks (Weinstock et al. 2016) Recycling of selection markers, like antibiotic resistances, as well as the removal of incorporated genes becomes easy and reliable.
Also interesting for future protein related applications is, that a putative secretion signal was identified (Weinstock et al. 2016)
Fun facts:
The epithet natriegens was chosen after the realization that salt is needed for it to grow, natrie stemming from the Latin word natrium meaning sodium and the Latin verb egere, meaning to need.
Aiyar, Sarah E, Tamas Gaal, and Richard L Gourse. 2002. RRNA Promoter Activity in the Fast-Growing Bacterium Vibrio Natriegens. Journal of bacteriology 184(5): 1349-58. http://www.ncbi.nlm.nih.gov/pubmed/11844764 (August 23, 2018).
Austin, Payne B, A Zachary, and R R Colwell. 2018. 28 International Association of Microbiological Societies Recognition of Beneckea Natriegens (Payne et al.; Baumann et al.) as a Member of the Genus Vibrio, as Previously Proposed by Webb and Payne . www.microbFiologyresearch.org (August 23, 2018).
Bi, Keran et al. 2016. Isolation and Molecular Identification of Vibrio Natriegens from Diseased Portunus Trituberculatus in China. Journal of the World Aquaculture Society 47(6): 854-61. http://doi.wiley.com/10.1111/jwas.12305 (September 29, 2018).
Carlson, Erik D, Rui Gan, C Eric Hodgman, and Michael C Jewett. 2012. Cell-Free Protein Synthesis: Applications Come of Age. Biotechnology advances 30(5): 1185–94. http://www.ncbi.nlm.nih.gov/pubmed/22008973 (October 1, 2018).
CHIEN, C et al. 2007. “Production of Poly-β-Hydroxybutyrate (PHB) by Vibrio Spp. Isolated from Marine Environment.” Journal of Biotechnology 132(3): 259–63. http://www.ncbi.nlm.nih.gov/pubmed/17416432 (September 22, 2018).
Coyer, J A, A Cabello-Pasini, H Swift, and R S Alberte. 1996. “N2 Fixation in Marine Heterotrophic Bacteria: Dynamics of Environmental and Molecular Regulation.” Proceedings of the National Academy of Sciences of the United States of America 93(8): 3575–80. http://www.ncbi.nlm.nih.gov/pubmed/11607653 (August 23, 2018).
Dalia, Triana N. et al. 2017. “Multiplex Genome Editing by Natural Transformation (MuGENT) for Synthetic Biology in Vibrio Natriegens .” ACS Synthetic Biology 6(9): 1650–55. http://www.ncbi.nlm.nih.gov/pubmed/28571309 (September 30, 2018).
Delpech, Roger. 2001. “Using Vibrio Natriegetis for Studying Bacterial Population Growth, Artificial Selection, and the Effects of UV Radiation and Photo-Reactivation.” Journal of Biological Education 35(2): 93–97. http://www.tandfonline.com/doi/abs/10.1080/00219266.2000.9655749 (August 23, 2018).
EAGON, R G. 1962. “Pseudomonas Natriegens, a Marine Bacterium with a Generation Time of Less than 10 Minutes.” Journal of bacteriology 83(4): 736–37. http://www.ncbi.nlm.nih.gov/pubmed/13888946 (August 23, 2018).
Failmezger, Jurek, Steffen Scholz, Bastian Blombach, and Martin Siemann-Herzberg. 2018. “Cell-Free Protein Synthesis From Fast-Growing Vibrio Natriegens.” Frontiers in Microbiology 9: 1146. https://www.frontiersin.org/article/10.3389/fmicb.2018.01146/full (September 30, 2018).
Fernández-Llamosas, Helga et al. 2017. “Speeding up Bioproduction of Selenium Nanoparticles by Using Vibrio Natriegens as Microbial Factory.” Scientific Reports 7(1): 16046. http://www.nature.com/articles/s41598-017-16252-1 (September 28, 2018).
Hoffart, Eugenia et al. 2017. “High Substrate Uptake Rates Empower Vibrio Natriegens as Production Host for Industrial Biotechnology.” Applied and environmental microbiology : AEM.01614-17. http://www.ncbi.nlm.nih.gov/pubmed/28887417 (October 6, 2018).
Lee, Henry H et al. 2016. “Vibrio Natriegens, a New Genomic Powerhouse.” bioRxiv : 058487. https://www.biorxiv.org/content/early/2016/06/12/058487 (September 30, 2018).
Lee, Henry H, Nili Ostrov, Michaela A Gold, and George M Church. 2017. “Recombineering in Vibrio Natriegens.” bioRxiv : 130088. https://www.biorxiv.org/content/early/2017/04/26/130088 (September 30, 2018).
Long, Christopher P., Jacqueline E. Gonzalez, Robert M. Cipolla, and Maciek R. Antoniewicz. 2017. “Metabolism of the Fast-Growing Bacterium Vibrio Natriegens Elucidated by 13C Metabolic Flux Analysis.” Metabolic Engineering 44: 191–97. http://www.ncbi.nlm.nih.gov/pubmed/29042298 (October 5, 2018).
Maida, I. et al. 2013. “Draft Genome Sequence of the Fast-Growing Bacterium Vibrio Natriegens Strain DSMZ 759.” Genome Announcements 1(4). http://www.ncbi.nlm.nih.gov/pubmed/23969053 (August 23, 2018).
Mehbub, Mohammad F. et al. 2018. “A Controlled Aquarium System and Approach to Study the Role of Sponge-Bacteria Interactions Using Aplysilla Rosea and Vibrio Natriegens.” Scientific Reports 8(1): 11801. http://www.nature.com/articles/s41598-018-30295-y (September 29, 2018).
Miura, A et al. 1981. “Growth-Rate-Dependent Regulation of Ribosome Synthesis in E. Coli: Expression of the LacZ and GalK Genes Fused to Ribosomal Promoters.” Cell 25(3): 773–82. http://www.ncbi.nlm.nih.gov/pubmed/6793240 (September 22, 2018).
Mullenger, L., and Nijole R. Gill. 1973. “Vibrio Natriegens: A Rapidly Growing Micro-Organism Ideally Suited for Class Experiments.” Journal of Biological Education 7(5): 33–39. http://www.tandfonline.com/doi/abs/10.1080/00219266.1973.9653881 (August 21, 2018).
Nazly, E D. 1980. 116 Journal of General Microbiology Adenine Nucleotide Pools During Starvation of Beneckea Natviegens . www.microbiologyresearch.org (October 7, 2018).
Nissen, Hilde, Mikal Heldal, and Svein Norland. 1987. “Growth, Elemental Composition, and Formation of Polyphosphate Bodies in Vibrio Natriegens Cultures Shifted from Phosphate-Limited to Phosphate-Pulsed Media.” Canadian Journal of Microbiology 33(7): 583–88. http://www.nrcresearchpress.com/doi/10.1139/m87-101 (September 22, 2018).
Payne, W J, R G Eagon, and A K Williams. 1960. SOME OBSERVATIONS ON THE PHYSIOLOGY OF PSEUDOMONAS NATRIEGENS NOV. SPEC. 1) . https://link.springer.com/content/pdf/10.1007/BF02538432.pdf (August 21, 2018).
Payne, William J. STUDIES ON BACTERIAL UTILIZATION OF URONIC ACIDS III. INDUCTION OF OXIDATIVE ENZYMES IN A MARINE ISOLATE’ . https://www.ncbi.nlm.nih.gov/pmc/articles/PMC290205/pdf/jbacter00506-0093.pdf (August 21, 2018).
Pollack-Berti, Amber, Michael S. Wollenberg, and Edward G. Ruby. 2010. “Natural Transformation of Vibrio Fischeri Requires TfoX and TfoY.” Environmental Microbiology 12(8): no-no. http://doi.wiley.com/10.1111/j.1462-2920.2010.02250.x (September 30, 2018).
Simons, Keryn L., Susan M. Thomas, and Peter A. Anderson. 2010. “Identification of Vibrio Natriegens UvrA and UvrB Genes and Analysis of Gene Regulation Using Transcriptional Reporter Plasmids.” The Journal of Microbiology 48(5): 644–56. http://www.ncbi.nlm.nih.gov/pubmed/21046343 (September 22, 2018).
Des Soye, Benjamin J. et al. 2018. “Establishing a High-Yielding Cell-Free Protein Synthesis Platform Derived from Vibrio Natriegens .” ACS Synthetic Biology 7(9): 2245–55. http://www.ncbi.nlm.nih.gov/pubmed/30107122 (September 30, 2018).
Webb, C D, and W J Payne. 1971. “Influence of Na+ on Synthesis of Macromolecules by a Marine Bacterium.” Applied microbiology 21(6): 1080–88. http://www.ncbi.nlm.nih.gov/pubmed/4327612 (August 23, 2018).
Weinstock, Matthew T, Eric D Hesek, Christopher M Wilson, and Daniel G Gibson. 2016. “Vibrio Natriegens as a Fast-Growing Host for Molecular Biology.” Nature Methods 13(10): 849–51. http://www.ncbi.nlm.nih.gov/pubmed/27571549 (September 29, 2018).
Wiegand, Daniel J., Henry H. Lee, Nili Ostrov, and George M. Church. 2018. “Establishing a Cell-Free Vibrio Natriegens Expression System.” ACS Synthetic Biology : acssynbio.8b00222. http://pubs.acs.org/doi/10.1021/acssynbio.8b00222 (September 30, 2018).
Zachary, A. 1976. “Physiology and Ecology of Bacteriophages of the Marine Bacterium Beneckea Natriegens: Salinity.” Applied and environmental microbiology 31(3): 415–22. http://www.ncbi.nlm.nih.gov/pubmed/938035 (August 23, 2018).
Zachary, Arthur. 1974. 27 APPLIED MICROBIOLOGY NOTES Isolation of Bacteriophages of the Marine Bacterium Beneckea Natriegens from Coastal Salt Marshes’ . https://www.ncbi.nlm.nih.gov/pmc/articles/PMC380186/pdf/applmicro00040-0188.pdf (September 28, 2018).
Zacheray, A. 1978. An Ecological Study of Bacteriophages of Vibrio Natriegens . Canadian Journal of Microbiology 24(3): 321-24. http://www.nrcresearchpress.com/doi/10.1139/m78-053 (September 23, 2018).
Brazelton, VA et al. 2016. A quick guide to CRISPR sgRNA design tools. GM Crops Food . 2015; 6(4): 266–276. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5033207/pdf/kgmc-06-04-1137690.pdf
Nishimasu, H., Ran, F. A., Hsu, P. D., Konermann, S., Shehata, S., Dohmae, N., Ishitani, R., Zhang, F., Nureki, O. (2014) Crystal Structure of Cas9 in Complex with Guide RNA and Target DNA, Cell , 156 (5), 935-949. https://www.cell.com/action/showPdf?pii=S0092-8674%2814%2900156-1
Jinek, M., Jiang, F., Taylor, D. W., Sternberg, S. H., Kaya, E., Ma, E., Anders, C., Hauer, M., Zhou, K., Lin, S., Kaplan, M., Iavarone, A. T., Charpentier, E., Nogale, E., Doudna, J. A. (2014) Structures of Cas9 Endonucleases Reveal RNA-Mediated Conformational Activation, Science , 343 (6176), 1-28. http://science.sciencemag.org/content/343/6176/1247997
Deltcheva, E., Chylinski, K., Sharma, C. M., Gonzales, K., Chao, Y., Pirzada, Z. A., Eckert, M. R., Vogel, J., Charpentier, E. (2011) CRISPR RNA maturation by trans -encoded small RNA and host factor RNase III, Nature , 471, 602-609. https://www.nature.com/articles/nature09886
Gasiunas, R. Barrangou, P. Horvath, V. Siksnys. 2012. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria.
Proc. Natl. Acad. Sci. USA , 109 (2012), pp. E2579-E2586. http://www.pnas.org/content/pnas/109/39/E2579.full.pdf
Jinek, K. Chylinski, I. Fonfara, M. Hauer, J.A. Doudna, E. Charpentier. 2012. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science , 337 (2012), pp. 816-821. http://science.sciencemag.org/content/337/6096/816
Doudna, J. A. & Charpentier, E. 2014.Genome editing. The new frontier of genome engineering with CRISPR-Cas9 . Science 346, 1258096 (2014). http://science.sciencemag.org/content/346/6213/1258096.full
Charpentier, E. & Marraffini, L. A. 2014. Harnessing CRISPR-Cas9 immunity for genetic engineering . Curr. Opin. Microbiol. 19, 114–119 (2014). https://www.sciencedirect.com/science/article/pii/S1369527414000885?via%3Dihub
Hsu, P. D., Lander, E. S., Zhang, F. (2014) Development and Applications of CRISPR-Cas9 for Genome Engineering, Cell , 157, 1262-1276. https://www.cell.com/action/showPdf?pii=S0092-8674%2814%2900604-7
Mazhar Adli. 2018. The CRISPR tool kit for genome editing and beyond. Nature Communications volume 9, Article number: 1911 (2018) https://www.nature.com/articles/s41467-018-04252-2.pdf
Jansen, R., van Embden, J. D. A., Gaastra, W., Schouls, L. M. (2002) Identification of genes that are associated with DNA repeats in prokaryotes, Molecular Microbiology , 43 (6), 1565-1575. https://onlinelibrary.wiley.com/doi/epdf/10.1046/j.1365-2958.2002.02839.x
Ishino, Y., Shinagawa, H., Makino, K., Amemura, M., Nakata, A. (1987) Nucleotide Sequence of the iap Gene, Responsible for Alkaline Phosphatase Isozyme Conversion in Escherichia coli , and Identification of the Gene Product, Journal of Bacteriology , 169 (12), 5429-5433. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC213968/pdf/jbacter00202-0107.pdf
Qi et al. 2013. Repurposing CRISPR as an RNA-Guided Platform for Sequence-Specific Control of Gene Expression. Cell. 2013 Feb 28; 152(5): 1173–1183. https://www.cell.com/action/showPdf?pii=S0092-8674%2813%2900211-0
Barrangou, R., Fremaux, C., Deveau, H., Richards, M., Boyaval, P., Moineau, S., Romero, D. A., Horvath, P. (2007) CRISPR Provides Acquired Resistance Against Viruses in Prokaryotes,
Science
, 315, 1709-1712. http://science.sciencemag.org/content/315/5819/1709
Brazelton, VA et al. 2016. A quick guide to CRISPR sgRNA design tools. GM Crops Food . 2015; 6(4): 266–276. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5033207/pdf/kgmc-06-04-1137690.pdf
Nishimasu, H., Ran, F. A., Hsu, P. D., Konermann, S., Shehata, S., Dohmae, N., Ishitani, R., Zhang, F., Nureki, O. (2014) Crystal Structure of Cas9 in Complex with Guide RNA and Target DNA, Cell , 156 (5), 935-949. https://www.cell.com/action/showPdf?pii=S0092-8674%2814%2900156-1
Jinek, M., Jiang, F., Taylor, D. W., Sternberg, S. H., Kaya, E., Ma, E., Anders, C., Hauer, M., Zhou, K., Lin, S., Kaplan, M., Iavarone, A. T., Charpentier, E., Nogale, E., Doudna, J. A. (2014) Structures of Cas9 Endonucleases Reveal RNA-Mediated Conformational Activation, Science , 343 (6176), 1-28. http://science.sciencemag.org/content/343/6176/1247997
Deltcheva, E., Chylinski, K., Sharma, C. M., Gonzales, K., Chao, Y., Pirzada, Z. A., Eckert, M. R., Vogel, J., Charpentier, E. (2011) CRISPR RNA maturation by trans -encoded small RNA and host factor RNase III, Nature , 471, 602-609. https://www.nature.com/articles/nature09886
Proc. Natl. Acad. Sci. USA , 109 (2012), pp. E2579-E2586. http://www.pnas.org/content/pnas/109/39/E2579.full.pdf
Jinek, K. Chylinski, I. Fonfara, M. Hauer, J.A. Doudna, E. Charpentier. 2012. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science , 337 (2012), pp. 816-821. http://science.sciencemag.org/content/337/6096/816
Doudna, J. A. & Charpentier, E. 2014.Genome editing. The new frontier of genome engineering with CRISPR-Cas9 . Science 346, 1258096 (2014). http://science.sciencemag.org/content/346/6213/1258096.full
Charpentier, E. & Marraffini, L. A. 2014. Harnessing CRISPR-Cas9 immunity for genetic engineering . Curr. Opin. Microbiol. 19, 114–119 (2014). https://www.sciencedirect.com/science/article/pii/S1369527414000885?via%3Dihub
Hsu, P. D., Lander, E. S., Zhang, F. (2014) Development and Applications of CRISPR-Cas9 for Genome Engineering, Cell , 157, 1262-1276. https://www.cell.com/action/showPdf?pii=S0092-8674%2814%2900604-7
Mazhar Adli. 2018. The CRISPR tool kit for genome editing and beyond. Nature Communications volume 9, Article number: 1911 (2018) https://www.nature.com/articles/s41467-018-04252-2.pdf
Jansen, R., van Embden, J. D. A., Gaastra, W., Schouls, L. M. (2002) Identification of genes that are associated with DNA repeats in prokaryotes, Molecular Microbiology , 43 (6), 1565-1575. https://onlinelibrary.wiley.com/doi/epdf/10.1046/j.1365-2958.2002.02839.x
Ishino, Y., Shinagawa, H., Makino, K., Amemura, M., Nakata, A. (1987) Nucleotide Sequence of the iap Gene, Responsible for Alkaline Phosphatase Isozyme Conversion in Escherichia coli , and Identification of the Gene Product, Journal of Bacteriology , 169 (12), 5429-5433. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC213968/pdf/jbacter00202-0107.pdf
Qi et al. 2013. Repurposing CRISPR as an RNA-Guided Platform for Sequence-Specific Control of Gene Expression. Cell. 2013 Feb 28; 152(5): 1173–1183. https://www.cell.com/action/showPdf?pii=S0092-8674%2813%2900211-0
Barrangou, R., Fremaux, C., Deveau, H., Richards, M., Boyaval, P., Moineau, S., Romero, D. A., Horvath, P. (2007) CRISPR Provides Acquired Resistance Against Viruses in Prokaryotes,
Science
, 315, 1709-1712. http://science.sciencemag.org/content/315/5819/1709
Alber, B.E., Fuchs, G., 2002. Propionyl-coenzyme a synthase from Chloroflexus aurantiacus, a key enzyme of the 3-hydroxypropionate cycle for autotrophic CO2 fixation. J. Biol. Chem. 277, 12137–12143.
Gande, R., Dover, L.G., Krumbach, K., Besra, G.S., Sahm, H., Oikawa, T., Eggeling, L., 2007. The two carboxylases of corynebacterium glutamicum essential for fatty acid and mycolic acid synthesis. J. Bacteriol. 189, 5257–5264.
Hanko, E.K.R., Minton, N.P., Malys, N., 2017. Characterisation of a 3-hydroxypropionic acid-inducible system from Pseudomonas putida for orthogonal gene expression control in Escherichia coli and Cupriavidus necator. Sci. Rep. 7, 1724.
Janßen, H.J., Steinbüchel, A., 2014. Fatty acid synthesis in Escherichia coli and its applications towards the production of fatty acid based biofuels. Biotechnol Biofuels 7, 7.
Kildegaard, K.R., Jensen, N.B., Schneider, K., Czarnotta, E., Özdemir, E., Klein, T., Maury, J., Ebert, B.E., Christensen, H.B., Chen, Y., Kim, I.K., Herrgård, M.J., Blank, L.M., Forster, J., Nielsen, J., Borodina, I., 2016. Engineering and systems-level analysis of Saccharomyces cerevisiae for production of 3-hydroxypropionic acid via malonyl-CoA reductase-dependent pathway. Microb. Cell Fact. 15.
Lee, C.-K., Cheong, H.-K., Ryu, K.-S., Lee, J. Il, Lee, W., Jeon, Y.H., Cheong, C., 2008. Biotinoyl domain of human acetyl-CoA carboxylase: Structural insights into the carboxyl transfer mechanism. Proteins Struct. Funct. Bioinforma. 72, 613–624.
Liu, C., Ding, Y., Xian, M., Liu, M., Liu, H., Ma, Q., Zhao, G., 2017. Malonyl-CoA pathway: a promising route for 3-hydroxypropionate biosynthesis. Crit. Rev. Biotechnol. 37, 933–941.
Liu, C., Wang, Q., Xian, M., Ding, Y., Zhao, G., 2013. Dissection of Malonyl-Coenzyme A Reductase of Chloroflexus aurantiacus Results in Enzyme Activity Improvement. PLoS One 8, 1–8.
Raj, S.M., Rathnasingh, C., Jo, J.E., Park, S., 2008. Production of 3-hydroxypropionic acid from glycerol by a novel recombinant Escherichia coli BL21 strain. Process Biochem. 43, 1440–1446.
Rathnasingh, C., Raj, S.M., Lee, Y., Catherine, C., Ashok, S., Park, S., 2012. Production of 3-hydroxypropionic acid via malonyl-CoA pathway using recombinant Escherichia coli strains. J. Biotechnol. 157, 633–640.
Ro, D.K., Paradise, E.M., Quellet, M., Fisher, K.J., Newman, K.L., Ndungu, J.M., Ho, K.A., Eachus, R.A., Ham, T.S., Kirby, J., Chang, M.C.Y., Withers, S.T., Shiba, Y., Sarpong, R., Keasling, J.D., 2006. Production of the antimalarial drug precursor artemisinic acid in engineered yeast. Nature 440, 940–943.
Stephanopoulos, G., 2012. Synthetic biology and metabolic engineering. ACS Synth. Biol. 1, 514–525.
Stephanopoulos, G., 2007. Challenges in engineering microbes for biofuels production. Science 315, 801–4.
Valdehuesa, K.N.G., Liu, H., Nisola, G.M., Chung, W.J., Lee, S.H., Park, S.J., 2013. Recent advances in the metabolic engineering of microorganisms for the production of 3-hydroxypropionic acid as C3 platform chemical. Appl. Microbiol. Biotechnol. 97, 3309–3321.
Vidra, A., Németh, Á., 2017. Bio-based 3-hydroxypropionic Acid : A Review. period. Polytech. Chem. Eng. 1–11.
Werpy, T., Petersen, G., 2004. Top Value Added Chemicals from Biomass Volume I — Results of Screening for Potential Candidates from Sugars and Synthesis Gas Top Value Added Chemicals From Biomass Volume I : Results of Screening for Potential Candidates. Other Inf. PBD 1 Aug 2004 Medium: ED; Size: 76 pp. pages.

