| (28 intermediate revisions by 4 users not shown) | |||
| Line 5: | Line 5: | ||
<!--Banner--> | <!--Banner--> | ||
| − | + | <div class="container-fluid parallax" style="background-image: url('https://static.igem.org/mediawiki/2018/a/a4/T--Toulouse-INSA-UPS--project--Brice--Banner.jpg');background-size: cover; background-position-y: center; " id="BANNER"></div> | |
</html> {{Template:Toulouse-INSA-UPS/MENU}} <html> | </html> {{Template:Toulouse-INSA-UPS/MENU}} <html> | ||
| Line 17: | Line 17: | ||
<h1 class="heavy">EXPERIMENTS</h1> | <h1 class="heavy">EXPERIMENTS</h1> | ||
<hr/> | <hr/> | ||
| − | <p> | + | <p> Here, you will find all the necessary information about the wetlab materials and methods we used in the course of our project.</p> |
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
<!-- Menu Generalissime --> | <!-- Menu Generalissime --> | ||
<div class="accordion" id="accordionExperiment"> | <div class="accordion" id="accordionExperiment"> | ||
| Line 64: | Line 59: | ||
<h3><em>Escherichia coli</em> vectors</h3> | <h3><em>Escherichia coli</em> vectors</h3> | ||
<h4>pET28</h4> | <h4>pET28</h4> | ||
| − | <p>The pET28 vector contains two His Tags in | + | <p>The pET28 vector contains two His Tags in its MCS and is specified for protein expression at high yield. It gives the strain a resistance to Kanamycin and its replication origin is the pBR322 which is a mid - low copy number plasmid (~10). The vector used was a kind gift from IBCG library. </p> |
| + | <div class="center"> | ||
<figure class="figure"> | <figure class="figure"> | ||
<img style="width : 50%; heigth = auto;" src="https://static.igem.org/mediawiki/2018/4/40/T--Toulouse-INSA-UPS--Experiments--Youn--pET28a.PNG" class="figure-img img-fluid rounded" alt="A generic square placeholder image with rounded corners in a figure."> | <img style="width : 50%; heigth = auto;" src="https://static.igem.org/mediawiki/2018/4/40/T--Toulouse-INSA-UPS--Experiments--Youn--pET28a.PNG" class="figure-img img-fluid rounded" alt="A generic square placeholder image with rounded corners in a figure."> | ||
<figcaption class="figure-caption">Figure 1. Map of the pET28 with the MCS Highligthed</figcaption> | <figcaption class="figure-caption">Figure 1. Map of the pET28 with the MCS Highligthed</figcaption> | ||
| − | </figure> | + | </figure></div> |
<h4>pETDuet-1</h4> | <h4>pETDuet-1</h4> | ||
| − | <p>The pETDuet-1 is, | + | <p>The pETDuet-1 is, like the pET28, specialized in high yield protein production. Its main difference is that it contains two MCS and can therefore express two proteins at the same time under the same expression conditions. It bears ampicilin resistance and is a mid - low copy number plasmid. The plasmid used was a gift from IBCG library.</p> |
| + | <div class="center"> | ||
<figure class="figure"> | <figure class="figure"> | ||
<img style="width : 50%; heigth = auto;" src="https://static.igem.org/mediawiki/2018/d/d1/T--Toulouse-INSA-UPS--Experiments--Youn--pETDuet1.PNG" class="figure-img img-fluid rounded" alt="A generic square placeholder image with rounded corners in a figure."> | <img style="width : 50%; heigth = auto;" src="https://static.igem.org/mediawiki/2018/d/d1/T--Toulouse-INSA-UPS--Experiments--Youn--pETDuet1.PNG" class="figure-img img-fluid rounded" alt="A generic square placeholder image with rounded corners in a figure."> | ||
| − | <figcaption class="figure-caption">Figure | + | <figcaption class="figure-caption">Figure 2. Map of the pETDuet-1</figcaption> |
| − | </figure> | + | </figure></div> |
<h3><em>Pichia pastoris</em> vectors</h3> | <h3><em>Pichia pastoris</em> vectors</h3> | ||
| − | <h4> | + | <h4>pPICZ alpha</h4> |
| − | <p>The pPICZalpha | + | <p>The pPICZalpha is an integrative plasmid for <i>Pichia pastoris</i> protein expression and suitable for <i>E. coli</i> replication. Its MCS is suitable for protein expession using the methanol inductible AOX1 Pomoter (PAOX1) and offering the alpha factor secretion signal for protein purification from culture supernatant. It contains a His tag and a Myc Tag for affinity purification stategies.</p> |
| + | <div class="center"> | ||
<figure class="figure"> | <figure class="figure"> | ||
<img style="width : 50%; heigth = auto;" src="https://static.igem.org/mediawiki/2018/8/8e/T--Toulouse-INSA-UPS--Experiments--Youn--pPICZalpha.PNG" class="figure-img img-fluid rounded" alt="A generic square placeholder image with rounded corners in a figure."> | <img style="width : 50%; heigth = auto;" src="https://static.igem.org/mediawiki/2018/8/8e/T--Toulouse-INSA-UPS--Experiments--Youn--pPICZalpha.PNG" class="figure-img img-fluid rounded" alt="A generic square placeholder image with rounded corners in a figure."> | ||
| − | <figcaption class="figure-caption">Figure | + | <figcaption class="figure-caption">Figure 3. Map of the pPICZ alpha</figcaption> |
| − | </figure> | + | </figure></div> |
| − | <h4> | + | <h4>pGAPZalpha </h4> |
| − | <p>The pGAPZalpha is roughly the same as the pPICZalpha with | + | <p>The pGAPZalpha is roughly the same as the pPICZalpha with a different promoter that is the GAP Promoter, a glucose inductible promoter.</p> |
| + | <div class="center"> | ||
<figure class="figure"> | <figure class="figure"> | ||
<img style="width : 50%; heigth = auto;" src="https://static.igem.org/mediawiki/2018/9/97/T--Toulouse-INSA-UPS--Experiments--Youn--pGAPZalpha.PNG" class="figure-img img-fluid rounded" alt="A generic square placeholder image with rounded corners in a figure."> | <img style="width : 50%; heigth = auto;" src="https://static.igem.org/mediawiki/2018/9/97/T--Toulouse-INSA-UPS--Experiments--Youn--pGAPZalpha.PNG" class="figure-img img-fluid rounded" alt="A generic square placeholder image with rounded corners in a figure."> | ||
| − | <figcaption class="figure-caption">Figure | + | <figcaption class="figure-caption">Figure 4. Map of the pGAPZ alpha</figcaption> |
| − | <h3>Primer used</h3> | + | </figure></div> |
| − | + | <h3>Primer used</h3> | |
| − | + | <ul> | |
| + | <li>Cerberus Forward : TAAGAAGGAGATATACCATGGCGGAAGCGGGTATCACC</li> | ||
| + | <li>Cerberus Reverse : CTCGAGTGCGGCCGCAAGCTTCGGATCGTCCTATGATGGAGG </li> | ||
| + | <li>Sirius Forward: TAAGAAGGAGATATACCATGAATGCTACGCCAACTAAGGGTGC</li> | ||
| + | <li>Sirius Reverse: CTCGAGTGCGGCCGCAAGCTTAGCACCGGTGGAGTGACG</li> | ||
| + | <li>BirA Forward: AAGGAGATATACATATGAAGGATAACACCGTGCCACTGA</li> | ||
| + | <li>BirA Reverse: CTTTACCAGACTCGATTATTTTTCTGCACTACGCAGGGA</li> | ||
| + | <li>BFP Forward: ACCACAGCCAGGATCCTATGAGCGAACTGATCAAAGAGAACA</li> | ||
| + | <li>BFP Reverse: ATGCGGCCGCAAGCTTCTCATGCCATTCAATTTTCTGTGCT</li> | ||
| + | <li>RFP Forward: AGGAGATATACCATGGCTTCCTCCGAAGACGTTATCAAAG</li> | ||
| + | <li>RFP Reverse: GTGCGGCCGCAAGCTTAGCACCGGTGGAGTGACG</li> | ||
| + | <li>Scygonadin Forward: GGCTGAAGCTGAATTCGGCCAGGCACTCAACAAAC</li> | ||
| + | <li>Scygonadin Reverse: TGGGCCACGTGAATTCTCACTCATGCCATTCAATTTTCTG</li> | ||
| + | </ul> | ||
| + | |||
</div> | </div> | ||
</div> | </div> | ||
| Line 100: | Line 113: | ||
<h5 class="mb-0"> | <h5 class="mb-0"> | ||
| − | <h2> | + | <h2>Microbiological</h2> |
</h5> | </h5> | ||
| Line 110: | Line 123: | ||
<h3><em>Escherichia coli</em> strains</h3> | <h3><em>Escherichia coli</em> strains</h3> | ||
| − | Plasmid amplification strains | + | <strong>Plasmid amplification strains</strong> |
<ul> | <ul> | ||
| − | <li><em>Stellar</em></li> | + | <li><em>Stellar</em> F-, endA1, supE44, thi-1, recA1, relA1, gyrA96, phoA, Φ80d lacZΔ M15, Δ(lacZYA-argF) U169, Δ(mrr-hsdRMS-mcrBC), ΔmcrA, λ- </li> |
| − | <li><em>Top10</em></li> | + | <li><em>Top10</em> F- mcrA Δ(mrr-hsdRMS-mcrBC) φ80lacZΔM15 ΔlacX74 nupG recA1 araD139 Δ(ara-leu)7697 galE15 galK16 rpsL(Str<sup>R</sup>) endA1 λ<sup>-</sup> </li> |
</ul> | </ul> | ||
| − | Protein production strains | + | <strong>Protein production strains</strong> |
<ul> | <ul> | ||
| − | <li><em> | + | <li><em>BL21(DE3)</em> B F<sup>-</sup><em>ompT gal dcm lon hsdS<sub>B</sub></em>(<em>r<sub>B</sub><sup>-</sup>m<sub>B</sub><sup>-</sup></em>) λ(DE3[<em>lacI lacUV5-T7p07 ind1 sam7 nin5</em>])[<em>malB<sup>+</sup></em>]<sub>K-12</sub>(λ<sup>S</sup>)</li> |
| − | <li><em> | + | <li><em>Tuner</em> F<sup>-</sup> ompT hsdS<sub>B</sub> (r<sub>B</sub><sup>-</sup> m<sub>B</sub><sup>-</sup>) gal dcm lacY1(DE3)</li> |
</ul> | </ul> | ||
<h3><em>Pichia pastoris</em> strains</h3> | <h3><em>Pichia pastoris</em> strains</h3> | ||
| − | <p>For protein production we used GS200 Strain for Cerberus and X33 for Scygonadin</p> | + | <p>For protein production we used GS200(ΔARG4, ΔHIS4) Strain for Cerberus production and X33(wt) for Scygonadin</p> |
| + | <h3><em>Gluconacetobacter hansenii</em> strain ATCC 53582 </h3> | ||
</div> | </div> | ||
</div> | </div> | ||
| Line 141: | Line 155: | ||
<div id="collapseThreeInt" class="collapse " aria-labelledby="headingInternalThree" data-parent="#accordionInternal"> | <div id="collapseThreeInt" class="collapse " aria-labelledby="headingInternalThree" data-parent="#accordionInternal"> | ||
<div class="card-body"> | <div class="card-body"> | ||
| − | Click reagants | + | <h3 class="heavy">Click reagants</h3> |
<ul> | <ul> | ||
| − | <li>DBCO-Biotin</li> | + | <li><strong>DBCO-Biotin</strong>, Sigma CAS : 1255942-07-4</li> |
| − | <li>DBCO-Fluorescein</li> | + | <li><strong>DBCO-Fluorescein</strong>, Jena Bioscience Cat. No. : CLK-051-1</li> |
| − | <li>DBCO-Magnetic beads</li> | + | <li><strong>DBCO-Magnetic beads</strong>, Jena Bioscience Cat. No. : CLK-1037-1</li> |
| + | <li><strong>Fluorescein-Azide</strong>, Jena Bioscience Cat. No. : CLK-80101-5</li> | ||
| + | <li><strong>4-L-azidophenylanaline</strong>, Sigma Ref. : 06162</li> | ||
| + | |||
| + | </ul> | ||
| + | <h3 class="heavy">Graphene Functionalization</h3> | ||
| + | <ul> | ||
| + | <li><strong>Graphene</strong>, Sigma Ref. : 900561</li> | ||
| + | <li><strong>4-Ethynylaniline</strong>, Sigma Ref. : 481122</li> | ||
| + | <li><strong>Isopentyl Nitrite</strong>, Sigma Ref. : 150495</li> | ||
| + | <li><strong>N,N-Dimethylformamide</strong>, Sigma Ref. : 227056</li> | ||
| + | </ul> | ||
| + | <h3 class="heavy">Protein purification</h3> | ||
| + | <ul> | ||
| + | <li><strong>Cobalt Affinity Gel</strong>, Sigma Ref. : H8162</li> | ||
| + | <li><strong>Protease inhibitor cocktail tablet</strong>, Sigma Ref. : S8830</li> | ||
| + | </ul> | ||
| + | <h3 class="heavy">Cellulose Binding assay</h3> | ||
| + | <ul> | ||
| + | <li><strong>Avicel</strong>, Sigma Ref. : 11365 </li> | ||
</ul> | </ul> | ||
</div> | </div> | ||
| Line 154: | Line 187: | ||
</div> | </div> | ||
</div> | </div> | ||
| − | + | <br/> | |
<!-- Menu Methodes --> | <!-- Menu Methodes --> | ||
<div class="card"> | <div class="card"> | ||
| Line 192: | Line 225: | ||
<div id="collapseOneInt2" class="collapse " aria-labelledby="headingInternal2One" data-parent="#accordionInternal2"> | <div id="collapseOneInt2" class="collapse " aria-labelledby="headingInternal2One" data-parent="#accordionInternal2"> | ||
<div class="card-body"> | <div class="card-body"> | ||
| − | < | + | <h2 id="medium" class="heavy">Media</h2> |
<hr> | <hr> | ||
<p>To prepare each medium, autoclave 20 minutes at 121°C.</p> | <p>To prepare each medium, autoclave 20 minutes at 121°C.</p> | ||
| − | < | + | <h3 id="lb_medium" class="heavy"> LB Medium </h3><hr/> |
<table> | <table> | ||
<thead> | <thead> | ||
| Line 225: | Line 258: | ||
</table><p>** Only for solid medium.</p> | </table><p>** Only for solid medium.</p> | ||
| − | <p><em>Nb</em>: For culture with zeocin as an antibiotic, the NaCl concentration | + | <p><em>Nb</em>: For culture with zeocin as an antibiotic, the NaCl concentration has to be 4g/L.</p> |
| − | < | + | <h3 id="hs-medium" class="heavy">HS Medium</h3><hr/> |
<hr> | <hr> | ||
| Line 233: | Line 266: | ||
<tr> | <tr> | ||
<th align="center">Component</th> | <th align="center">Component</th> | ||
| − | |||
<th>Concentration (g/L)</th> | <th>Concentration (g/L)</th> | ||
</tr> | </tr> | ||
| Line 240: | Line 272: | ||
<tr> | <tr> | ||
<td align="center">Glucose</td> | <td align="center">Glucose</td> | ||
| − | |||
<td>20g</td> | <td>20g</td> | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
<td align="center">Yeast extract</td> | <td align="center">Yeast extract</td> | ||
| − | |||
<td>5g</td> | <td>5g</td> | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
<td align="center">Peptone</td> | <td align="center">Peptone</td> | ||
| − | |||
<td>5g</td> | <td>5g</td> | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
<td align="center">Na2HPO4</td> | <td align="center">Na2HPO4</td> | ||
| − | |||
<td>2.7g</td> | <td>2.7g</td> | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
<td align="center">Citric acid</td> | <td align="center">Citric acid</td> | ||
| − | |||
<td>1.5g</td> | <td>1.5g</td> | ||
</tr> | </tr> | ||
<tr> | <tr> | ||
<td align="center">**Agar</td> | <td align="center">**Agar</td> | ||
| − | |||
<td>15g</td> | <td>15g</td> | ||
</tr> | </tr> | ||
| Line 274: | Line 300: | ||
Autoclaving glucose separately from amino acids avoids Maillard reaction, which can result in the formation of toxic byproducts in the media.</p> | Autoclaving glucose separately from amino acids avoids Maillard reaction, which can result in the formation of toxic byproducts in the media.</p> | ||
| − | < | + | <h3 id="ypd_medium" class="heavy"> YPD Medium </h3><hr/> |
<table> | <table> | ||
<thead> | <thead> | ||
| Line 301: | Line 327: | ||
</tbody> | </tbody> | ||
</table><p>** Only for solid medium</p> | </table><p>** Only for solid medium</p> | ||
| − | <p><em>NB</em>: Autoclave | + | <p><em>NB</em>: Autoclave the glucose solution separately from the rest.<br> |
Autoclaving glucose separately from amino acids avoids Maillard reaction, which can result in the formation of toxic byproducts in the media.</p> | Autoclaving glucose separately from amino acids avoids Maillard reaction, which can result in the formation of toxic byproducts in the media.</p> | ||
| − | < | + | <h3 id="tb-medium">TB medium</h3><hr/> |
<hr> | <hr> | ||
| Line 328: | Line 354: | ||
</tbody> | </tbody> | ||
| − | </table><p>For solid media, add 15g of agarose | + | </table><p>For solid media, add 15g of agarose per liter.<br> |
Autoclave : 20min at 121°C</p> | Autoclave : 20min at 121°C</p> | ||
| − | < | + | <h3 id="md_medium" class="heavy">Minimal Dextrose Medium (MD)</h3><hr/> |
<p><strong>Medium for selection of recombinant Arg+ clones For 1 L MD agar plates you need</strong>:</p> | <p><strong>Medium for selection of recombinant Arg+ clones For 1 L MD agar plates you need</strong>:</p> | ||
| Line 345: | Line 371: | ||
<li>Dissolve 69 g YNB (without aminoacids; with ammonium sulfat) in 500 mL bidest water and filter sterilize.</li> | <li>Dissolve 69 g YNB (without aminoacids; with ammonium sulfat) in 500 mL bidest water and filter sterilize.</li> | ||
<li>Store at 4 °C.</li> | <li>Store at 4 °C.</li> | ||
| − | <li> | + | <li>Will last for one year.</li> |
</ul> | </ul> | ||
<p><strong>Biotin 500X stock solution</strong>:</p> | <p><strong>Biotin 500X stock solution</strong>:</p> | ||
| Line 353: | Line 379: | ||
<li>Durable for one year.</li> | <li>Durable for one year.</li> | ||
</ul> | </ul> | ||
| − | < | + | <h3 id="bmmy-medium" class="heavy">BMMY medium</h3> |
<hr> | <hr> | ||
<p>BMMY Buffered Methal-complex medium</p> | <p>BMMY Buffered Methal-complex medium</p> | ||
| Line 391: | Line 417: | ||
</tbody> | </tbody> | ||
</table><p><em>Storage conditions</em> : 4°C</p> | </table><p><em>Storage conditions</em> : 4°C</p> | ||
| − | < | + | <h3 id="bmgy-medium" class="heavy">BMGY medium</h3> |
<hr> | <hr> | ||
<p>BMGY Buffered Glycerol-complex medium</p> | <p>BMGY Buffered Glycerol-complex medium</p> | ||
| Line 420: | Line 446: | ||
<tr> | <tr> | ||
<td align="center">Glycerol</td> | <td align="center">Glycerol</td> | ||
| − | <td>1 | + | <td>1%</td> |
</tr> | </tr> | ||
<tr> | <tr> | ||
| Line 430: | Line 456: | ||
<h2 id="buffer">Buffer</h2> | <h2 id="buffer">Buffer</h2> | ||
<hr> | <hr> | ||
| − | < | + | <h3 id="materials" class="heavy">Materials</h3> |
<hr> | <hr> | ||
<ul> | <ul> | ||
| Line 439: | Line 465: | ||
<li>Acetic acid</li> | <li>Acetic acid</li> | ||
</ul> | </ul> | ||
| − | < | + | <h3 id="buffer-for-transformation" class="heavy">Buffer for transformation</h3> |
<hr> | <hr> | ||
| − | <h4 id="yetm-ph-7.5" class="heavy">YETM pH 7.5</h4> | + | <h4 id="yetm-ph-7.5" class="heavy">YETM pH 7.5</h4><hr/> |
<table> | <table> | ||
<thead> | <thead> | ||
| Line 469: | Line 495: | ||
</table><p>** Only for solid medium</p> | </table><p>** Only for solid medium</p> | ||
<p>Adjust to pH to 7.5 with kOH.</p> | <p>Adjust to pH to 7.5 with kOH.</p> | ||
| − | <h4 id="tfb1-ph-5.8" class="heavy">TFB1 pH 5.8</h4> | + | <h4 id="tfb1-ph-5.8" class="heavy">TFB1 pH 5.8</h4><hr/> |
<table> | <table> | ||
<thead> | <thead> | ||
| Line 500: | Line 526: | ||
</tbody> | </tbody> | ||
</table><p>Adjust to pH 5.8 with 0.2M acetic acid. Add dionized water up to 1L. Filter sterilize. Store refrigerated at 4°C.</p> | </table><p>Adjust to pH 5.8 with 0.2M acetic acid. Add dionized water up to 1L. Filter sterilize. Store refrigerated at 4°C.</p> | ||
| − | <h4 id="tfb2-ph-6.5" class="heavy">TFB2 pH 6.5</h4> | + | <h4 id="tfb2-ph-6.5" class="heavy">TFB2 pH 6.5</h4><hr/> |
<table> | <table> | ||
<thead> | <thead> | ||
| Line 527: | Line 553: | ||
</tbody> | </tbody> | ||
</table><p>Adjust to pH 6.5 with kOH. Add dionized water to 1L. Filter sterilize. Store refrigerated at 4°C.</p> | </table><p>Adjust to pH 6.5 with kOH. Add dionized water to 1L. Filter sterilize. Store refrigerated at 4°C.</p> | ||
| − | <h4 id="guanidine-hcl-6m-ph-7.5" class="heavy">Guanidine HCl 6M pH 7.5</h4> | + | <h4 id="guanidine-hcl-6m-ph-7.5" class="heavy">Guanidine HCl 6M pH 7.5</h4><hr/> |
<table> | <table> | ||
<thead> | <thead> | ||
| Line 549: | Line 575: | ||
<h2 id="culture-conditions" class="heavy">Culture conditions</h2> | <h2 id="culture-conditions" class="heavy">Culture conditions</h2> | ||
<hr> | <hr> | ||
| − | < | + | <h3 class="heavy"><em>Escherichia coli</em></h3> |
<ul> | <ul> | ||
<li>37°C</li> | <li>37°C</li> | ||
| Line 557: | Line 583: | ||
</li> | </li> | ||
</ul> | </ul> | ||
| − | < | + | <h3 class="heavy"><em>Pichia pastoris</em></h3> |
<ul> | <ul> | ||
<li>37 °C</li> | <li>37 °C</li> | ||
| Line 565: | Line 591: | ||
</li> | </li> | ||
</ul> | </ul> | ||
| − | < | + | <h3><em>Gluconacetobacter hansenii</em></h3> |
<ul> | <ul> | ||
<li>30°C</li> | <li>30°C</li> | ||
| Line 583: | Line 609: | ||
<h5 class="mb-0"> | <h5 class="mb-0"> | ||
| − | <h2>Cloning & | + | <h2>Cloning & Transformation</h2> |
</h5> | </h5> | ||
| Line 591: | Line 617: | ||
<div id="collapseTwoInt2" class="collapse " aria-labelledby="headingInternal2Two" data-parent="#accordionInternal2"> | <div id="collapseTwoInt2" class="collapse " aria-labelledby="headingInternal2Two" data-parent="#accordionInternal2"> | ||
<div class="card-body"> | <div class="card-body"> | ||
| − | < | + | <h2 id="clonage">Cloning</h2> |
<hr> | <hr> | ||
| − | < | + | <h3 id="insert-preparation" class="heavy">1 . Insert preparation</h3> |
<hr> | <hr> | ||
| Line 635: | Line 661: | ||
<li>Cycle 3 : - 72°C 2 min</li> | <li>Cycle 3 : - 72°C 2 min</li> | ||
</ul> | </ul> | ||
| − | < | + | <h3 id="vector-preparation" class="heavy">2 . Vector preparation</h3> |
<hr> | <hr> | ||
| − | + | <h5 id="miniprep-and-midiprep" class="heavy">2 . 1 . Miniprep and Midiprep</h5> | |
| − | <h5 id="miniprep-and-midiprep" class="heavy">2. Miniprep and Midiprep</h5> | + | |
<ul> | <ul> | ||
<li><em><strong>Miniprep</strong></em></li> | <li><em><strong>Miniprep</strong></em></li> | ||
</ul> | </ul> | ||
| − | |||
This protocol is extracted from the GenElute™ Plasmid Miniprep Kit from Sigma Aldricht.</p> | This protocol is extracted from the GenElute™ Plasmid Miniprep Kit from Sigma Aldricht.</p> | ||
<ol> | <ol> | ||
| Line 648: | Line 672: | ||
- High-copy plasmid : 1-3 ml<br> | - High-copy plasmid : 1-3 ml<br> | ||
- Low-copy plasmid : 1-5 ml<br> | - Low-copy plasmid : 1-5 ml<br> | ||
| − | Pellet 1-5ml of an overnight recombinant E.coli culture by | + | Pellet 1-5ml of an overnight recombinant <i>E. coli</i> culture by centrifuging at <strong>4,500rpm</strong> for <strong>4min</strong>. Discard the supernatant.</li> |
<li><em>Resuspend cells</em><br> | <li><em>Resuspend cells</em><br> | ||
| − | Completely resuspend the bacterial pellet with <strong> | + | Completely resuspend the bacterial pellet with <strong>200 µl</strong> of the Resuspension Solution. Vortex or pipette up and down to thoroughly mix the cells until homogeneous.</li> |
<li><em>Lyse cells</em><br> | <li><em>Lyse cells</em><br> | ||
| − | Lyse the resuspended cells by adding <strong> | + | Lyse the resuspended cells by adding <strong>200 µl</strong> of the lysis Solution. Immediately mix the contents by gentle inversion (6-8 times) until the mixture becomes clear and viscous.<br> |
<em>NB</em> : Do not allow the lysis reaction to exceed 5 minutes.</li> | <em>NB</em> : Do not allow the lysis reaction to exceed 5 minutes.</li> | ||
<li><em>Neutralize</em><br> | <li><em>Neutralize</em><br> | ||
| Line 659: | Line 683: | ||
<li><em>Prepare column</em><br> | <li><em>Prepare column</em><br> | ||
Insert a GenElute Miniprep Binding Column into a provided microcentrifuge tube.<br> | Insert a GenElute Miniprep Binding Column into a provided microcentrifuge tube.<br> | ||
| − | Add <strong> | + | Add <strong>500 µl</strong> of the Column Preparation Solution to each miniprep column and centrifuge <strong>12,000 x g</strong> during <strong>1min</strong>.</li> |
<li><em>Load cleared lysate</em><br> | <li><em>Load cleared lysate</em><br> | ||
Transfer the cleared lysate from step 4 to the column prepared in step 5 and centrifuge at <strong>12,000 x g</strong> for <strong>1 min</strong>. Discard the flow-through liquid.<br> | Transfer the cleared lysate from step 4 to the column prepared in step 5 and centrifuge at <strong>12,000 x g</strong> for <strong>1 min</strong>. Discard the flow-through liquid.<br> | ||
| Line 677: | Line 701: | ||
<li><strong>Midiprep</strong></li> | <li><strong>Midiprep</strong></li> | ||
</ul> | </ul> | ||
| − | + | ||
<p>This protocol is extracted from the QIAfilter™ Plasmid Midi Kit (25) from QIAGEN.</p> | <p>This protocol is extracted from the QIAfilter™ Plasmid Midi Kit (25) from QIAGEN.</p> | ||
<p>High-copy plasmid : 25 ml<br> | <p>High-copy plasmid : 25 ml<br> | ||
| Line 696: | Line 720: | ||
<li>Air-dry the pellet for 20 min and redissolve DNA in 150 µl miliQ water</li> | <li>Air-dry the pellet for 20 min and redissolve DNA in 150 µl miliQ water</li> | ||
</ol> | </ol> | ||
| − | <h5 id="preparative-digest-gel-extraction" class="heavy">3. Preparative digest gel extraction</h5> | + | <h5 id="preparative-digest-gel-extraction" class="heavy">2 . 3 . Preparative digest gel extraction</h5> |
<hr> | <hr> | ||
| − | |||
<table> | <table> | ||
<thead> | <thead> | ||
| Line 725: | Line 748: | ||
</tbody> | </tbody> | ||
</table><p>** If you have 2 enzymes 0.5µl of each ones.</p> | </table><p>** If you have 2 enzymes 0.5µl of each ones.</p> | ||
| − | <h4 id="gel-extraction-purification" class="heavy">3. Gel extraction purification</h4> | + | <h4 id="gel-extraction-purification" class="heavy">3 . Gel extraction purification</h4> |
<hr> | <hr> | ||
<p>This protocol is extracted from the NucleoSpin® kit. We would extract both vector and insert with this protocol.</p> | <p>This protocol is extracted from the NucleoSpin® kit. We would extract both vector and insert with this protocol.</p> | ||
| Line 731: | Line 754: | ||
<li>Excise DNA fragment and weight it.</li> | <li>Excise DNA fragment and weight it.</li> | ||
<li>Solubilize gel slice : for each 100 mg of agarose gel, add 200 µl of Buffer NTI. Incubate sample for 10 min at 50°C, mix by inverting the tube every 2-3 min.</li> | <li>Solubilize gel slice : for each 100 mg of agarose gel, add 200 µl of Buffer NTI. Incubate sample for 10 min at 50°C, mix by inverting the tube every 2-3 min.</li> | ||
| − | <li>Bind DNA : place a column (provided in the kit) into a collection tube (2 ml, also provided) and load up to 700µl sample. Centrifuge for <strong>30s at 11 000g</strong>. Discard flow-through and place the column back into the collection tube.</li> | + | <li>Bind DNA : place a column (provided in the kit) into a collection tube (2 ml, also provided) and load up to 700µl sample. Centrifuge for <strong>30s at 11,000g</strong>. Discard flow-through and place the column back into the collection tube.</li> |
| − | <li>Wash silica membrane : <strong>add 700µl Buffer NT3</strong> to the column. Centrifuge for <strong>30s at 11 000g</strong>. Discard flow-through and place the column back into the collection tube. <strong>Repeat this step</strong> one more time.</li> | + | <li>Wash silica membrane : <strong>add 700µl Buffer NT3</strong> to the column. Centrifuge for <strong>30s at 11,000g</strong>. Discard flow-through and place the column back into the collection tube. <strong>Repeat this step</strong> one more time.</li> |
| − | <li>Dry silica membrane : centrifuge <strong>1 min at 11 000g</strong> to remove Buffer NT3 completely.</li> | + | <li>Dry silica membrane : centrifuge <strong>1 min at 11,000g</strong> to remove Buffer NT3 completely.</li> |
| − | <li>Elute DNA : place the column into a new 1.5ml Eppendorf. <strong>Add | + | <li>Elute DNA : place the column into a new 1.5ml Eppendorf. <strong>Add 20 µl of H2O MQ</strong> and incubate at room temperature (18 - 25°C) for 1 min. Centrifuge for <strong>1 min at 11,000g</strong>.</li> |
</ol> | </ol> | ||
| − | <h4 id="in-fusion" class="heavy">4. | + | <h4 id="in-fusion" class="heavy">4 . InFusion cloning</h4> |
<hr> | <hr> | ||
| − | <h5 id="clonage-1" class="heavy">1. | + | <h5 id="clonage-1" class="heavy">4 . 1 . Cloning</h5> |
<p><strong>Materials</strong></p> | <p><strong>Materials</strong></p> | ||
<ol> | <ol> | ||
| Line 773: | Line 796: | ||
</tbody> | </tbody> | ||
</table> | </table> | ||
| − | <a href="#transformation_for_in_fusion" class="int_link" class="heavy"></a><h5> 2. Transformation | + | <a href="#transformation_for_in_fusion" class="int_link" class="heavy"></a><h5>4 . 2. Transformation into Stellar® competent cells </h5> |
| − | < | + | <h2 id="transformation" class="heavy">Transformation</h2> |
<hr> | <hr> | ||
| − | <h3 id="escherichia--coli" class="heavy">1. Escherichia coli</h3> | + | <h3 id="escherichia--coli" class="heavy">1 . <em>Escherichia coli</em></h3><hr/> |
| − | <h4 id="preparation-of-competent-cells--bl21" class="heavy">1. Preparation of competent cells : BL21</h4> | + | <h4 id="preparation-of-competent-cells--bl21" class="heavy">1 . 1 . Preparation of competent cells : BL21(DE3)</h4> |
<hr> | <hr> | ||
<p><strong>Protocol</strong></p> | <p><strong>Protocol</strong></p> | ||
| Line 797: | Line 820: | ||
immediately on dry ice. Stire cells frozen at -80°C.</li> | immediately on dry ice. Stire cells frozen at -80°C.</li> | ||
</ol> | </ol> | ||
| − | <h4 id="h4-idtransformation_of_competent_cells-2.-transformation-of-competent-cells-h4" class="heavy"></h4><h4 id="transformation_of_competent_cells" class="heavy"> 2. Transformation of Competent Cells </h4> | + | <h4 id="h4-idtransformation_of_competent_cells-2.-transformation-of-competent-cells-h4" class="heavy"></h4><h4 id="transformation_of_competent_cells" class="heavy">1 . 2 . Transformation of Competent Cells </h4> |
<hr> | <hr> | ||
<p><strong>Protocol</strong></p> | <p><strong>Protocol</strong></p> | ||
| Line 823: | Line 846: | ||
<li>Transform 100 μL of cells with 1 μL (10 pg) of pUC19 monomer (0.01 μg/μL).</li> | <li>Transform 100 μL of cells with 1 μL (10 pg) of pUC19 monomer (0.01 μg/μL).</li> | ||
<li>Plate 0.25 mL of transformation mixture. Incubate overnight at 37°C.</li> | <li>Plate 0.25 mL of transformation mixture. Incubate overnight at 37°C.</li> | ||
| − | <li>Count CFU and calculate efficiency. Efficiency | + | <li>Count CFU and calculate efficiency. Efficiency of colonies per μg of colonies 4 * 10<sup>5</sup>. You should<br> |
| − | obtain 1-5 X | + | obtain 1-5 X 10<sup>7</sup>/μg<sub>DNA</sub> from competent cells after one freeze-thaw cycle.</li> |
</ol> | </ol> | ||
| − | <h3 id="pichia-pastoris" class="heavy">2. Pichia | + | <h3 id="pichia-pastoris" class="heavy">2 . <i>Pichia pastoris</i></h3> |
<hr> | <hr> | ||
| − | <h4 id="electroporation" class="heavy">1. Electroporation</h4> | + | <h4 id="electroporation" class="heavy">2 . 1 . Electroporation</h4> |
<hr> | <hr> | ||
| − | <h5 id="competent-cells-preparation" class="heavy">1. | + | <h5 id="competent-cells-preparation" class="heavy">2 . 1 . 1 . Competent cells preparation</h5> |
<p><strong>Materials</strong><br> | <p><strong>Materials</strong><br> | ||
- YPD/0.02M HEPES buffer<br> | - YPD/0.02M HEPES buffer<br> | ||
| Line 836: | Line 859: | ||
- 1.0M Sorbitol<br> | - 1.0M Sorbitol<br> | ||
- 1.5ml tubes<br> | - 1.5ml tubes<br> | ||
| − | - | + | - Sterile water (1.250l)<br> |
- Centrifuge (with temperature control)</p> | - Centrifuge (with temperature control)</p> | ||
<p><strong>Protocol</strong></p> | <p><strong>Protocol</strong></p> | ||
<ol> | <ol> | ||
| − | <li>Grow 500ml cells to DO 1.3 to 1.5 at 600 nm and <strong>keep in | + | <li>Grow 500ml cells to DO 1.3 to 1.5 at 600 nm and <strong>keep in ice for the duration of the procedure)</strong></li> |
<li>Centrifuge cells for 5 min (all centrifugation steps were at 4000g at 8°C)</li> | <li>Centrifuge cells for 5 min (all centrifugation steps were at 4000g at 8°C)</li> | ||
<li>Add 100ml YPD/0.02M HEPES</li> | <li>Add 100ml YPD/0.02M HEPES</li> | ||
| Line 857: | Line 880: | ||
<li>Place in -80°C freezer until needed</li> | <li>Place in -80°C freezer until needed</li> | ||
</ol> | </ol> | ||
| − | <h5 id="electroporation-pichia-pastoris" class="heavy">2. Electroporation Pichia | + | <h5 id="electroporation-pichia-pastoris" class="heavy">2 . 1 . 2 Electroporation <i>Pichia pastoris</i></h5> |
<hr> | <hr> | ||
<p><strong>Materials:</strong><br> | <p><strong>Materials:</strong><br> | ||
| Line 877: | Line 900: | ||
</ol> | </ol> | ||
<p>It can take around one week to have one colony on plate.</p> | <p>It can take around one week to have one colony on plate.</p> | ||
| − | <h4 id="heat-shock-pichia-pastoris" class="heavy">2. Heat shock Pichia | + | <h4 id="heat-shock-pichia-pastoris" class="heavy">2 . 2 . Heat shock <i>Pichia pastoris</i></h4> |
<hr> | <hr> | ||
| − | <h5 id="cell-preparation" class="heavy">1. Cell Preparation</h5> | + | <h5 id="cell-preparation" class="heavy">2 . 2 . 1 . Cell Preparation</h5> |
<p><strong>Protocols</strong></p> | <p><strong>Protocols</strong></p> | ||
<ol> | <ol> | ||
| Line 891: | Line 914: | ||
</ol> | </ol> | ||
<p>BEDS solution is composed of 10 mM bicine-NaOH, pH 8.3, 3% (v/v) ethylene glycol, 5% (v/v) dimethyl sulfoxide (DMSO), and 1 M sorbitol.</p> | <p>BEDS solution is composed of 10 mM bicine-NaOH, pH 8.3, 3% (v/v) ethylene glycol, 5% (v/v) dimethyl sulfoxide (DMSO), and 1 M sorbitol.</p> | ||
| − | <h5 id="transformation-1" class="heavy">2. Transformation</h5> | + | <h5 id="transformation-1" class="heavy">2 . 2 . 2 Transformation</h5> |
<ol> | <ol> | ||
<li>Mix DNA with cells (50 µl cells, 5-8 µl DNA)</li> | <li>Mix DNA with cells (50 µl cells, 5-8 µl DNA)</li> | ||
| Line 902: | Line 925: | ||
<li>Plate</li> | <li>Plate</li> | ||
</ol> | </ol> | ||
| − | <h3 id="transformation_for_in_fusion" class="heavy"> 3. Transformation for | + | <h3 id="transformation_for_in_fusion" class="heavy"> 3 . Transformation for InFusion </h3> |
<h4 id="stellar®-competent" class="heavy">Stellar® competent</h4> | <h4 id="stellar®-competent" class="heavy">Stellar® competent</h4> | ||
<hr> | <hr> | ||
| Line 944: | Line 967: | ||
<h2 id="protein-production" class="heavy">Protein production</h2> | <h2 id="protein-production" class="heavy">Protein production</h2> | ||
<hr> | <hr> | ||
| − | <h3 id="protein-production---echerichia-coli" class="heavy">1. Protein production - <em> | + | <h3 id="protein-production---echerichia-coli" class="heavy">1. Protein production - <em>Escherichia coli</em></h3> |
<hr> | <hr> | ||
<p><strong>Material</strong></p> | <p><strong>Material</strong></p> | ||
| Line 958: | Line 981: | ||
<li>Day one : | <li>Day one : | ||
<ul> | <ul> | ||
| − | <li>Transform your plasmid in < | + | <li>Transform your plasmid in <em>E. coli BL21(DE3)</em> competent cells plate on LB Agar + Your favourite antibiotic and incubate O/N at 37°C</li> |
</ul> | </ul> | ||
</li> | </li> | ||
| Line 964: | Line 987: | ||
<ul> | <ul> | ||
<li>Pick 3 colonies and seed then in 9ml LB + Your favourite antibiotic. Incubate O/N at 37°C under 130 rpm shaking.</li> | <li>Pick 3 colonies and seed then in 9ml LB + Your favourite antibiotic. Incubate O/N at 37°C under 130 rpm shaking.</li> | ||
| − | <li>Autoclave 800ml LB 10 g/L NaCl in a 2L baffled flask and | + | <li>Autoclave 800ml LB 10 g/L NaCl in a 2L baffled flask and preheat it at 37°C O/N.</li> |
</ul> | </ul> | ||
</li> | </li> | ||
| Line 971: | Line 994: | ||
<li>Take a sample of the 800ml culture and measure OD600 as blank.</li> | <li>Take a sample of the 800ml culture and measure OD600 as blank.</li> | ||
<li>Take 1ml of preculture and measure OD600, seed the preculture in the 800ml LB and take OD.</li> | <li>Take 1ml of preculture and measure OD600, seed the preculture in the 800ml LB and take OD.</li> | ||
| − | <li>Calculate | + | <li>Calculate OD600 assuming that it doubles every 20 minutes and check the OD until it reaches 0.5 ~0.6. Once the exponontial phase is reached, induce with 800µl of 1M IPTG stock solution and incubate at 37°C for 4h or O/N at 16°C.</li> |
<li>Pellet cells by centrifugate at 5000 x g for 10 minutes and resuspend in 10ml TB 1x 10mM Imidazole if growth was performed at 37°C of 10ml of O/N.</li> | <li>Pellet cells by centrifugate at 5000 x g for 10 minutes and resuspend in 10ml TB 1x 10mM Imidazole if growth was performed at 37°C of 10ml of O/N.</li> | ||
<li>Freeze cells for later purification or wait at least 30 minutes to continue the protocol.</li> | <li>Freeze cells for later purification or wait at least 30 minutes to continue the protocol.</li> | ||
<li>Sonication : Run on cycle of 1 minute and one of 30 seconds, both at 25 of intensity.</li> | <li>Sonication : Run on cycle of 1 minute and one of 30 seconds, both at 25 of intensity.</li> | ||
| − | <li>Pellet cells by centrifugate at | + | <li>Pellet cells by centrifugate at 60,000 x g for 30 minutes and withdraw the supernatant that is the <strong>cell free extract</strong>.</li> |
| − | <li>Then continue to the | + | <li>Then continue to the protein purification protocol.</li> |
</ul> | </ul> | ||
</li> | </li> | ||
</ul> | </ul> | ||
| − | <h3 id="protein-production---pichia-pastoris">2. Protein production - <em>Pichia | + | <h3 id="protein-production---pichia-pastoris">2. Protein production - <em>Pichia pastoris</em></h3><hr/> |
| − | <p>Expression can be done in either | + | <p>Expression can be done in either YPD medium (1% yeast extract, 2% peptone, 2% glucose) or Yeast Nitrogen Base with 0.5% glucose.</p> |
<ol> | <ol> | ||
<li>Using a single colony, inoculate 10 ml of YPD. Grow at 28–30°C in a shaking incubator (250–300 rpm) O/N.</li> | <li>Using a single colony, inoculate 10 ml of YPD. Grow at 28–30°C in a shaking incubator (250–300 rpm) O/N.</li> | ||
| Line 994: | Line 1,017: | ||
<hr/> | <hr/> | ||
<h4 id="imac-column-purification" class="heavy">1 . 1 .IMAC column purification</h4> | <h4 id="imac-column-purification" class="heavy">1 . 1 .IMAC column purification</h4> | ||
| − | <p>The volumes will be expressed as CV ( | + | <p>The volumes will be expressed as CV (Column Volume) which refers to the volume of resin deposited.</p> |
<p><strong>Materials</strong></p> | <p><strong>Materials</strong></p> | ||
<ul> | <ul> | ||
| Line 1,028: | Line 1,051: | ||
<hr> | <hr> | ||
<h4 id="rac" class="heavy">2 . 1 . RAC preparation </h4><hr/> | <h4 id="rac" class="heavy">2 . 1 . RAC preparation </h4><hr/> | ||
| − | + | <p><strong>Materials</strong></p> | |
| − | + | <ul> | |
| − | + | <li>Cellulose</li> | |
| − | + | <li>H<sub>2</sub>PO<sub>3</sub></li> | |
| − | + | <li>Na<sub>2</sub>CO<sub>3</sub> 2M</li> | |
| − | + | <li>Deionized water</li> | |
| + | <li>Ice</li> | ||
| + | </ul> | ||
<p><strong>Protocol</strong></p> | <p><strong>Protocol</strong></p> | ||
<ol> | <ol> | ||
<li>Weigth about 0.2 g of Cellulose, put it in a 50ml Falcon tube and add 0.6 ml of H20 to wet it.</li> | <li>Weigth about 0.2 g of Cellulose, put it in a 50ml Falcon tube and add 0.6 ml of H20 to wet it.</li> | ||
| − | <li>Add 10ml of ice cold | + | <li>Add 10ml of ice cold H<sub>2</sub>PO<sub>3</sub> and vortex<br> |
incubate 1h on ice and vortex envery 20 minutes</li> | incubate 1h on ice and vortex envery 20 minutes</li> | ||
<li>Add 40 ml of ice cold H2O and vortex every 10ml | <li>Add 40 ml of ice cold H2O and vortex every 10ml | ||
| Line 1,044: | Line 1,069: | ||
</ul> | </ul> | ||
</li> | </li> | ||
| − | <li>Centrifugate at 10 000 x g for 20 minutes and discart supernatant.</li> | + | <li>Centrifugate at 10,000 x g for 20 minutes and discart supernatant.</li> |
| − | <li>Wash with 45ml of H20 and 0.5ml of | + | <li>Wash with 45ml of H20 and 0.5ml of Na<sub>2</sub>CO<sub>3</sub> 2M |
<ul> | <ul> | ||
<li>Repeat 4 times</li> | <li>Repeat 4 times</li> | ||
| Line 1,058: | Line 1,083: | ||
<li>Regenerated Amorphous Cellulose</li> | <li>Regenerated Amorphous Cellulose</li> | ||
<li>Ethylene Glycol</li> | <li>Ethylene Glycol</li> | ||
| − | <li | + | <li> RAC previously prepared</li> |
</ul> | </ul> | ||
<p><strong>Method</strong></p> | <p><strong>Method</strong></p> | ||
| Line 1,069: | Line 1,094: | ||
<li>Analyse by SDS PAGE.</li> | <li>Analyse by SDS PAGE.</li> | ||
</ul> | </ul> | ||
| − | <h3 id="protein-concentration-using-vivaspin-column" class="heavy">Protein concentration using vivaspin column</h3> | + | <h3 id="protein-concentration-using-vivaspin-column" class="heavy">3 .Protein concentration using vivaspin column</h3> |
<hr> | <hr> | ||
<p><strong>Materials</strong></p> | <p><strong>Materials</strong></p> | ||
| Line 1,077: | Line 1,102: | ||
<p><strong>Protocol</strong><br> | <p><strong>Protocol</strong><br> | ||
Add your favourite purified protein (Dont go over 20ml) and equilibrate with and empty tube willed with water (DO NOT DISCART AFTER) and centrifugate for 5 minutes at 4500 x g. Observe the eluted volume and adapt the time for the desired final volume (this is protein dependant).</p> | Add your favourite purified protein (Dont go over 20ml) and equilibrate with and empty tube willed with water (DO NOT DISCART AFTER) and centrifugate for 5 minutes at 4500 x g. Observe the eluted volume and adapt the time for the desired final volume (this is protein dependant).</p> | ||
| − | <h3 id="desalting-protein-using-pd-10-column" class="heavy">Desalting protein using PD-10 column</h3> | + | <h3 id="desalting-protein-using-pd-10-column" class="heavy">4 . Desalting protein using PD-10 column</h3> |
<hr> | <hr> | ||
<ol> | <ol> | ||
| Line 1,134: | Line 1,159: | ||
<li>Protein sample (concentration about 5mg/ml)</li> | <li>Protein sample (concentration about 5mg/ml)</li> | ||
<li>Ethylene glycol</li> | <li>Ethylene glycol</li> | ||
| − | <li | + | <li>RAC previously prepared</li> |
</ul> | </ul> | ||
<p><strong>Method</strong></p> | <p><strong>Method</strong></p> | ||
| Line 1,147: | Line 1,172: | ||
</li> | </li> | ||
<li>Incubate at room temperature for 30 minutes</li> | <li>Incubate at room temperature for 30 minutes</li> | ||
| − | <li>Centrifugate at >10 000 x g for 10 minutes | + | <li>Centrifugate at >10,000 x g for 10 minutes |
<ul> | <ul> | ||
<li>Collect supernatant fraction</li> | <li>Collect supernatant fraction</li> | ||
| Line 1,154: | Line 1,179: | ||
<li>Wash with 5 x RAC Volume of Tris HCl | <li>Wash with 5 x RAC Volume of Tris HCl | ||
<ul> | <ul> | ||
| − | <li>Centrifugate at >10 000 x g for 10 minutes</li> | + | <li>Centrifugate at >10,000 x g for 10 minutes</li> |
</ul> | </ul> | ||
</li> | </li> | ||
| Line 1,162: | Line 1,187: | ||
<h3 class="heavy">Strain Promoted Alkyne Azide Cycloaddition</h3> | <h3 class="heavy">Strain Promoted Alkyne Azide Cycloaddition</h3> | ||
<p><strong>Method</strong></p> | <p><strong>Method</strong></p> | ||
| − | <p>Add in 100& | + | <p>Add in 100μL a ratio of 1:10 of Azide:DBCO, cover with aluminium foil and incubate O/N under shaking</p> |
</div> | </div> | ||
</div> | </div> | ||
| Line 1,187: | Line 1,212: | ||
<div id="NAV_ICON_BAR" class="nav-left-col sticky-bottom d-none d-xl-block"> | <div id="NAV_ICON_BAR" class="nav-left-col sticky-bottom d-none d-xl-block"> | ||
<ul class="nav justify-content-center"> | <ul class="nav justify-content-center"> | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
<li class="nav-item"> | <li class="nav-item"> | ||
<!--Anchor to TOP--> | <!--Anchor to TOP--> | ||
| Line 1,199: | Line 1,219: | ||
</a> | </a> | ||
</li> | </li> | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
| − | + | ||
</ul> | </ul> | ||
</div> | </div> | ||
Latest revision as of 22:49, 17 October 2018
EXPERIMENTS
Here, you will find all the necessary information about the wetlab materials and methods we used in the course of our project.
Escherichia coli vectors
pET28
The pET28 vector contains two His Tags in its MCS and is specified for protein expression at high yield. It gives the strain a resistance to Kanamycin and its replication origin is the pBR322 which is a mid - low copy number plasmid (~10). The vector used was a kind gift from IBCG library.
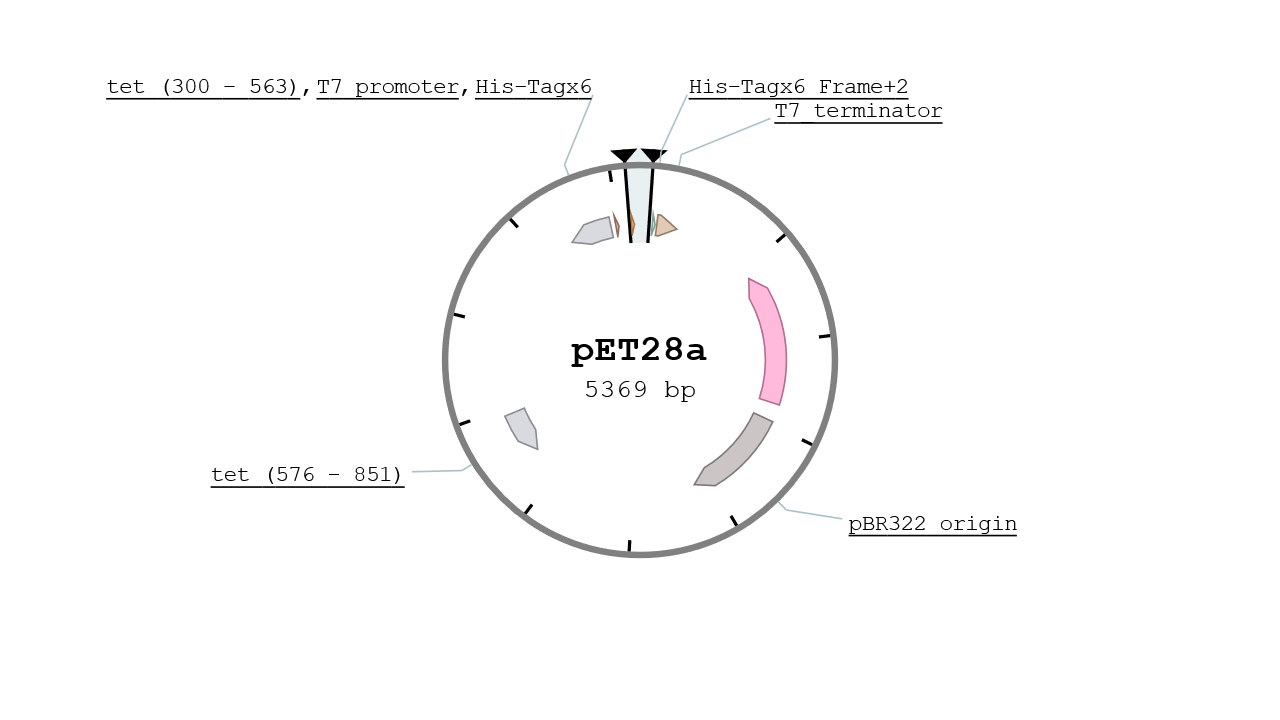
pETDuet-1
The pETDuet-1 is, like the pET28, specialized in high yield protein production. Its main difference is that it contains two MCS and can therefore express two proteins at the same time under the same expression conditions. It bears ampicilin resistance and is a mid - low copy number plasmid. The plasmid used was a gift from IBCG library.

Pichia pastoris vectors
pPICZ alpha
The pPICZalpha is an integrative plasmid for Pichia pastoris protein expression and suitable for E. coli replication. Its MCS is suitable for protein expession using the methanol inductible AOX1 Pomoter (PAOX1) and offering the alpha factor secretion signal for protein purification from culture supernatant. It contains a His tag and a Myc Tag for affinity purification stategies.

pGAPZalpha
The pGAPZalpha is roughly the same as the pPICZalpha with a different promoter that is the GAP Promoter, a glucose inductible promoter.

Primer used
- Cerberus Forward : TAAGAAGGAGATATACCATGGCGGAAGCGGGTATCACC
- Cerberus Reverse : CTCGAGTGCGGCCGCAAGCTTCGGATCGTCCTATGATGGAGG
- Sirius Forward: TAAGAAGGAGATATACCATGAATGCTACGCCAACTAAGGGTGC
- Sirius Reverse: CTCGAGTGCGGCCGCAAGCTTAGCACCGGTGGAGTGACG
- BirA Forward: AAGGAGATATACATATGAAGGATAACACCGTGCCACTGA
- BirA Reverse: CTTTACCAGACTCGATTATTTTTCTGCACTACGCAGGGA
- BFP Forward: ACCACAGCCAGGATCCTATGAGCGAACTGATCAAAGAGAACA
- BFP Reverse: ATGCGGCCGCAAGCTTCTCATGCCATTCAATTTTCTGTGCT
- RFP Forward: AGGAGATATACCATGGCTTCCTCCGAAGACGTTATCAAAG
- RFP Reverse: GTGCGGCCGCAAGCTTAGCACCGGTGGAGTGACG
- Scygonadin Forward: GGCTGAAGCTGAATTCGGCCAGGCACTCAACAAAC
- Scygonadin Reverse: TGGGCCACGTGAATTCTCACTCATGCCATTCAATTTTCTG
Escherichia coli strains
Plasmid amplification strains- Stellar F-, endA1, supE44, thi-1, recA1, relA1, gyrA96, phoA, Φ80d lacZΔ M15, Δ(lacZYA-argF) U169, Δ(mrr-hsdRMS-mcrBC), ΔmcrA, λ-
- Top10 F- mcrA Δ(mrr-hsdRMS-mcrBC) φ80lacZΔM15 ΔlacX74 nupG recA1 araD139 Δ(ara-leu)7697 galE15 galK16 rpsL(StrR) endA1 λ-
- BL21(DE3) B F-ompT gal dcm lon hsdSB(rB-mB-) λ(DE3[lacI lacUV5-T7p07 ind1 sam7 nin5])[malB+]K-12(λS)
- Tuner F- ompT hsdSB (rB- mB-) gal dcm lacY1(DE3)
Pichia pastoris strains
For protein production we used GS200(ΔARG4, ΔHIS4) Strain for Cerberus production and X33(wt) for Scygonadin
Gluconacetobacter hansenii strain ATCC 53582
Click reagants
- DBCO-Biotin, Sigma CAS : 1255942-07-4
- DBCO-Fluorescein, Jena Bioscience Cat. No. : CLK-051-1
- DBCO-Magnetic beads, Jena Bioscience Cat. No. : CLK-1037-1
- Fluorescein-Azide, Jena Bioscience Cat. No. : CLK-80101-5
- 4-L-azidophenylanaline, Sigma Ref. : 06162
Graphene Functionalization
- Graphene, Sigma Ref. : 900561
- 4-Ethynylaniline, Sigma Ref. : 481122
- Isopentyl Nitrite, Sigma Ref. : 150495
- N,N-Dimethylformamide, Sigma Ref. : 227056
Protein purification
- Cobalt Affinity Gel, Sigma Ref. : H8162
- Protease inhibitor cocktail tablet, Sigma Ref. : S8830
Cellulose Binding assay
- Avicel, Sigma Ref. : 11365
No dogs were harmed over the course of this iGEM project.
The whole Toulouse INSA-UPS team wants to thank our sponsors, especially:









And many more. For futher information about our sponsors, please consult our Sponsors page.
The content provided on this website is the fruit of the work of the Toulouse INSA-UPS iGEM Team. As a deliverable for the iGEM Competition, it falls under the Creative Commons Attribution 4.0. Thus, all content on this wiki is available under the Creative Commons Attribution 4.0 license (or any later version). For futher information, please consult the official website of Creative Commons.
This website was designed with Bootstrap (4.1.3). Bootstrap is a front-end library of component for html, css and javascript. It relies on both Popper and jQuery. For further information, please consult the official website of Bootstrap.

