Huiyee11186 (Talk | contribs) |
|||
| (39 intermediate revisions by 4 users not shown) | |||
| Line 6: | Line 6: | ||
<body data-spy="scroll" data-target=".navbar-example"> | <body data-spy="scroll" data-target=".navbar-example"> | ||
<div class="container content"> | <div class="container content"> | ||
| − | <h1 class="head">Design</h1> | + | <div class="headstyle"> |
| + | <h1 class="head">Design</h1> | ||
| + | </div> | ||
| + | <div class="righttitle"> | ||
| + | <h6 class="subtitle">Bring Solutions, Not Problems</h6> | ||
| + | </div> | ||
<div class="navbar-example"> | <div class="navbar-example"> | ||
<div class="row"> | <div class="row"> | ||
| Line 18: | Line 23: | ||
<a class="list-group-item list-group-item-action" href="#dual_plasmid_system">Dual plasmid system</a> | <a class="list-group-item list-group-item-action" href="#dual_plasmid_system">Dual plasmid system</a> | ||
<a class="list-group-item list-group-item-action" href="#pH_sensing_system">pH sensing system</a> | <a class="list-group-item list-group-item-action" href="#pH_sensing_system">pH sensing system</a> | ||
| − | <a class="list-group-item list-group-item-action" href="# | + | <a class="list-group-item list-group-item-action" href="#Reference">References</a> |
<a class="list-group-item list-group-item-action" href="#"><i class="fa fa-arrow-up fa-1x" aria-hidden="true"></i></a> | <a class="list-group-item list-group-item-action" href="#"><i class="fa fa-arrow-up fa-1x" aria-hidden="true"></i></a> | ||
</div> | </div> | ||
| Line 26: | Line 31: | ||
<div class="container"> | <div class="container"> | ||
<div id="Overview"> | <div id="Overview"> | ||
| − | <h3>Overview of Designed | + | <h3>Overview of Designed Pathway</h3> |
| + | <img class="gif" src="https://static.igem.org/mediawiki/2018/2/2c/T--NCKU_Tainan--wetdesign.gif" alt="Total pathway"> | ||
<p class="pcontent">Calvin-Benson cycle is one of the most important pathways for inorganic carbon to be converted into organic carbon in the carbon cycle. | <p class="pcontent">Calvin-Benson cycle is one of the most important pathways for inorganic carbon to be converted into organic carbon in the carbon cycle. | ||
Plant, algae, and cyanobacteria utilize light as energy source for Calvin-Benson cycle. | Plant, algae, and cyanobacteria utilize light as energy source for Calvin-Benson cycle. | ||
Taking the advantage of the pentose phosphate pathway, a native metabolic pathway of <i>E. coli</i>, | Taking the advantage of the pentose phosphate pathway, a native metabolic pathway of <i>E. coli</i>, | ||
| − | only two additional enzymes will be needed to reconstruct the pathway in <i>E. coli</i> -- PRK and | + | only two additional enzymes will be needed to reconstruct the pathway in <i>E. coli</i> -- PRK and Rubisco, |
which we will describe more in detail. | which we will describe more in detail. | ||
The primary product of the pathway is pyruvate, which can be utilized to produce various valuable products. | The primary product of the pathway is pyruvate, which can be utilized to produce various valuable products. | ||
| Line 44: | Line 50: | ||
W3110 (K-12 laboratory strain) is reported to be resilient in a stressed environment. | W3110 (K-12 laboratory strain) is reported to be resilient in a stressed environment. | ||
We expected that W3110 will grow well even if the sole carbon source is xylose. | We expected that W3110 will grow well even if the sole carbon source is xylose. | ||
| − | W3110(L5T7) (provided by Dr. Ng) is a constructed lab strain based on W3110. | + | W3110 (L5T7) (provided by Dr. Ng) is a constructed lab strain based on W3110. |
T7 polymerase was inserted into its genome. | T7 polymerase was inserted into its genome. | ||
</p> | </p> | ||
| Line 59: | Line 65: | ||
</p> | </p> | ||
<img class="gif" src="https://static.igem.org/mediawiki/2018/7/71/T--NCKU_Tainan--design_PRK.gif" alt="PRK"> | <img class="gif" src="https://static.igem.org/mediawiki/2018/7/71/T--NCKU_Tainan--design_PRK.gif" alt="PRK"> | ||
| − | <p class="pcontent">Steps involved in expressing PRK in <i>E. coli.</i> | + | |
| − | + | <div class="row"> | |
| − | + | <a class="btn col-md-12" data-toggle="collapse" href="#PRK_how_to_construct" role="button" aria-expanded="false" aria-controls="multiCollapseExample1"> | |
| − | + | How do we construct this part? | |
| − | + | <i class="fa fa-arrow-down fa-10" aria-hidden="true"></i> | |
| − | + | </a> | |
| − | + | </div> | |
| − | + | <div class="collapse multi-collapse" id="PRK_how_to_construct"> | |
| + | <div class="card card-body"> | ||
| + | <p class="pcontent">Steps involved in expressing PRK in <i>E. coli.</i> | ||
| + | We initially confirm the gene sequence of <i>Synechococcus elongtus</i> <i>prk</i> from NCBI gene database. | ||
| + | We then codon optimized the sequence so <i>E. coli</i> can express the protein properly. | ||
| + | The optimized sequence was sent to IDT for gene synthesis. | ||
| + | We PCR amplified the gene fragments and digest it with restriction enzymes HindIII and SpeI. | ||
| + | After digestion, we ligate the fragments into pSB3K3 plasmid with P<sub>LacI</sub>-rbs (B0034) located upstream of the fragment. | ||
| + | The plasmid was then transformed into DH5 alpha. | ||
| + | </p> | ||
| + | </div> | ||
| + | </div> | ||
<img class="bigimg" src="https://static.igem.org/mediawiki/2018/d/dd/T--NCKU_Tainan--design_PRK_construction.png" alt="PRK construction picture"> | <img class="bigimg" src="https://static.igem.org/mediawiki/2018/d/dd/T--NCKU_Tainan--design_PRK_construction.png" alt="PRK construction picture"> | ||
| − | <h5 class="question"> | + | |
| + | <h5 class="question">How do we test its function?</h5> | ||
<p class="pcontent">We initially decided to measure the concentration of RuBP by HPLC. | <p class="pcontent">We initially decided to measure the concentration of RuBP by HPLC. | ||
Our instructors pointed out some difficulties in HPLC measurement such as excessive noise signal in our sample. | Our instructors pointed out some difficulties in HPLC measurement such as excessive noise signal in our sample. | ||
| Line 82: | Line 100: | ||
<p class="pcontent">Ribulose-1,5-biphosphate carboxylase/oxygenase is one of the world’s most abundant enzyme. | <p class="pcontent">Ribulose-1,5-biphosphate carboxylase/oxygenase is one of the world’s most abundant enzyme. | ||
It catalyzes the conversion of inorganic carbon into organic carbon. | It catalyzes the conversion of inorganic carbon into organic carbon. | ||
| − | In our designed pathway, the function of the | + | In our designed pathway, the function of the Rubisco is to convert ribulose-1,5-biphosphate (RuBP) from the upper pathway and carbon dioxide into 3-phosphoglycerate (3PGA). |
3PGA will then be converted to pyruvate by the native metabolic system of <i>E. coli</i>. | 3PGA will then be converted to pyruvate by the native metabolic system of <i>E. coli</i>. | ||
After mining information from various publications, | After mining information from various publications, | ||
| − | we selected | + | we selected Rubisco from <i>Synechococcus elongatus</i> PCC 7002, which is a well-studied cyanobacteria. |
Its genome is completely sequenced and it is often used as a model organism for gene manipulation. | Its genome is completely sequenced and it is often used as a model organism for gene manipulation. | ||
| − | Previous research has utilized <i>E. coli</i> as a host of random mutagenesis to enhance the activity of <i>Synechococcus</i> | + | Previous research has utilized <i>E. coli</i> as a host of random mutagenesis to enhance the activity of <i>Synechococcus</i> Rubisco. |
</p> | </p> | ||
<img class="gif" src="https://static.igem.org/mediawiki/2018/8/85/T--NCKU_Tainan--design_Rubisco.gif" alt="Rubisco"> | <img class="gif" src="https://static.igem.org/mediawiki/2018/8/85/T--NCKU_Tainan--design_Rubisco.gif" alt="Rubisco"> | ||
| − | < | + | |
| − | <p class="pcontent">Akin to the construction of | + | <div class="row"> |
| − | + | <a class="btn col-md-12" data-toggle="collapse" href="#RuBisCO_how_to_construct" role="button" aria-expanded="false" aria-controls="multiCollapseExample1"> | |
| − | + | How do we construct this part? | |
| − | + | <i class="fa fa-arrow-down fa-10" aria-hidden="true"></i> | |
| − | + | </a> | |
| − | + | </div> | |
| − | + | <div class="collapse multi-collapse" id="RuBisCO_how_to_construct"> | |
| − | + | <div class="card card-body"> | |
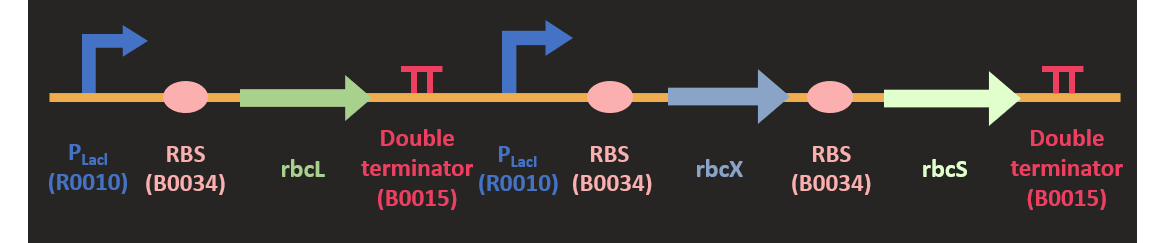
| + | <p class="pcontent">Akin to the construction of <i>prk</i>, we codon optimized the sequence of three <i>rbc</i> subunit and | ||
| + | clone it into pSB1C3 plasmid with HindIII and SpeI. | ||
| + | The sequence and the size of <i>rbcL</i> is much larger than other subunit, | ||
| + | so we separated <i>rbcL</i> from <i>rbcX</i> and <i>rbcS</i> subunits. <i>rbcX</i> and <i>rbcS</i> is separated by a <i>rbs</i> (B0034) for the convenience of construction. | ||
| + | We attached two different promoters upstream of the <i>rbc</i>. They are P<sub>LacI</sub> and P<sub>T7</sub> promoter. | ||
| + | Since we would like to increase the expression of this protein in the metabolic pathway, | ||
| + | we would like to test various promoter combination to find out the most efficient combination for our pathway. | ||
| + | </p> | ||
| + | </div> | ||
| + | </div> | ||
<img class="bigimg" src="https://static.igem.org/mediawiki/2018/a/ad/T--NCKU_Tainan--design_RBC_construction.png" alt="RBC construction picture"> | <img class="bigimg" src="https://static.igem.org/mediawiki/2018/a/ad/T--NCKU_Tainan--design_RBC_construction.png" alt="RBC construction picture"> | ||
<h5 class="question">How do we test its function?</h5> | <h5 class="question">How do we test its function?</h5> | ||
| − | <p class="pcontent">Measurement of 3PGA or pyruvate concentration could not directly reflect the activity of | + | <p class="pcontent">Measurement of 3PGA or pyruvate concentration could not directly reflect the activity of Rubisco |
since both of them are important metabolites that will flow to downstream metabolic pathway. | since both of them are important metabolites that will flow to downstream metabolic pathway. | ||
| − | We then decided to determine its function by total solution test which we will mention below. | + | We then decided to determine its function by a total solution test which we will mention below. |
</p> | </p> | ||
</div> | </div> | ||
| Line 109: | Line 137: | ||
<h3>CA</h3> | <h3>CA</h3> | ||
<h5 class="question">What is its function?</h5> | <h5 class="question">What is its function?</h5> | ||
| − | <p class="pcontent"> | + | <p class="pcontent">Rubisco is the rate-limiting enzyme in carbon fixation. |
Oxygen competes with CO<sub>2</sub> as a substrate for Rubisco, giving rise to photorespiration. | Oxygen competes with CO<sub>2</sub> as a substrate for Rubisco, giving rise to photorespiration. | ||
To overcome this problem, some photosynthetic organisms have evolved their own carbon concentrating | To overcome this problem, some photosynthetic organisms have evolved their own carbon concentrating | ||
| − | + | mechanism (CCM), which helps to maintain a sufficient amount of CO<sub>2</sub> around Rubisco. | |
</p> | </p> | ||
<p class="pcontent">We are inspired by the carbon concentrating mechanisms (CCM) of cyanobacteria. | <p class="pcontent">We are inspired by the carbon concentrating mechanisms (CCM) of cyanobacteria. | ||
In cyanobacteria, Rubisco and carbonic anhydrase (CA) is encapsulated in a microcompartment, the carboxysome. | In cyanobacteria, Rubisco and carbonic anhydrase (CA) is encapsulated in a microcompartment, the carboxysome. | ||
Carbonic anhydrase, also known as carbonate dehydratase, is involved in the interconversion between CO<sub>2</sub> and HCO<sub>3</sub><sup>-</sup>. This enzyme can be found in most organisms, including <i>E. coli</i> but the difference is its catalyzing rate in hydration and dehydration of CO2. Therefore, | Carbonic anhydrase, also known as carbonate dehydratase, is involved in the interconversion between CO<sub>2</sub> and HCO<sub>3</sub><sup>-</sup>. This enzyme can be found in most organisms, including <i>E. coli</i> but the difference is its catalyzing rate in hydration and dehydration of CO2. Therefore, | ||
| − | we will incorporate into our system the carbonic anhydrase gene from <i>Synechococcus elongatus | + | we will incorporate into our system the carbonic anhydrase gene from <i>Synechococcus elongatus</i> PCC 7002. |
</p> | </p> | ||
<img class="gif" src="https://static.igem.org/mediawiki/2018/3/34/T--NCKU_Tainan--design_CA.gif" alt="Rubisco"> | <img class="gif" src="https://static.igem.org/mediawiki/2018/3/34/T--NCKU_Tainan--design_CA.gif" alt="Rubisco"> | ||
| − | < | + | <div class="row"> |
| − | <p class="pcontent">We first codon optimized the sequence and insert it into the empty pSB1C3 plasmid with HindIII and | + | <a class="btn col-md-12" data-toggle="collapse" href="#CA_how_to_construct" role="button" aria-expanded="false" aria-controls="multiCollapseExample1"> |
| − | + | How do we construct this part? | |
| − | + | <i class="fa fa-arrow-down fa-10" aria-hidden="true"></i> | |
| − | + | </a> | |
| − | + | </div> | |
| + | <div class="collapse multi-collapse" id="CA_how_to_construct"> | ||
| + | <div class="card card-body"> | ||
| + | <p class="pcontent">We first codon optimized the sequence and insert it into the empty pSB1C3 plasmid with HindIII and | ||
| + | SpeI just as mentioned above. In our optimized sequence, we have already designed a P<sub>T7</sub> promoter | ||
| + | in front of <i>ca</i>, so we can directly ligate it into the plasmid. | ||
| + | The constructed basic part is then linked with other basic parts to complete our construction. | ||
| + | </p> | ||
| + | </div> | ||
| + | </div> | ||
<img class="bigimg" src="https://static.igem.org/mediawiki/2018/7/78/T--NCKU_Tainan--design_CA_construction.png" alt="CA Construction picture"> | <img class="bigimg" src="https://static.igem.org/mediawiki/2018/7/78/T--NCKU_Tainan--design_CA_construction.png" alt="CA Construction picture"> | ||
<h5 class="question">How do we test its function?</h5> | <h5 class="question">How do we test its function?</h5> | ||
<p class="pcontent">To measure the enzyme activity of CA, we compare the conversion rate of carbon dioxide to | <p class="pcontent">To measure the enzyme activity of CA, we compare the conversion rate of carbon dioxide to | ||
bicarbonate ion. After saturated CO<sub>2</sub> solution is prepared, we add fixed amount of bacteria broth that | bicarbonate ion. After saturated CO<sub>2</sub> solution is prepared, we add fixed amount of bacteria broth that | ||
| − | contains CA construction into the solution. We then measure the time | + | contains CA construction into the solution. We then measure the time taken for the pH value to decrease from 8.3 to 6.3. |
We compare the measured time interval with the time interval that enzyme was not added to determine the enzyme activity of CA. | We compare the measured time interval with the time interval that enzyme was not added to determine the enzyme activity of CA. | ||
</p> | </p> | ||
</div> | </div> | ||
<div id="dual_plasmid_system"> | <div id="dual_plasmid_system"> | ||
| − | <h3> | + | <h3>Dual Plasmid System</h3> |
<p class="pcontent">We decided to construct the whole pathway with the dual plasmid system. | <p class="pcontent">We decided to construct the whole pathway with the dual plasmid system. | ||
Previously, every basic part was the backbone conserved in the backbone of pSB1C3. | Previously, every basic part was the backbone conserved in the backbone of pSB1C3. | ||
We then link the construction together and | We then link the construction together and | ||
| − | even change the backbone of some composite parts to pSB3K3 for lower protein expression. | + | even change the backbone of some composite parts to pSB3K3 for a lower protein expression. |
</p> | </p> | ||
<h5 class="question">Rubisco whole protein in pSB1C3</h5> | <h5 class="question">Rubisco whole protein in pSB1C3</h5> | ||
| − | <p class="pcontent">We link each basic | + | <p class="pcontent">We link each basic part together with biobrick standard method. |
| − | We link P<sub>T7</sub>-rbcL and P<sub>T7</sub>-rbcX-rbcS together. | + | We link P<sub>T7</sub>-<i>rbcL</i> and P<sub>T7</sub>-<i>rbcX</i>-<i>rbcS</i> together. |
| − | The former, the insert, was digested with EcoRI and SpeI and the | + | The former, the insert, was digested with EcoRI and SpeI and the latter, the backbone, |
is digested with EcoRI and XbaI. | is digested with EcoRI and XbaI. | ||
We ligate the backbone with the insert to complete this composite part. | We ligate the backbone with the insert to complete this composite part. | ||
</p> | </p> | ||
| − | <h5 class="question"> | + | <h5 class="question"><i>prk</i> gene into pSB3K3</h5> |
| − | <p class="pcontent">PRK catalyzes the reaction of | + | <p class="pcontent">PRK catalyzes the reaction of converting Ru5P into RuBP. |
Not native to the host, RuBP is, nonetheless, toxic to <i>E. coli</i>. | Not native to the host, RuBP is, nonetheless, toxic to <i>E. coli</i>. | ||
| − | We hope that expression of PRK | + | We hope that the expression of PRK could be lower in the host so we change the backbone of it into pSB3K3. |
We selected J04450 from the distributed kit that under the backbone of pSB3K3, | We selected J04450 from the distributed kit that under the backbone of pSB3K3, | ||
which will express red color after the formation of the colony. We digest both backbone and insert with EcoRI and PstI and ligate both fragments. | which will express red color after the formation of the colony. We digest both backbone and insert with EcoRI and PstI and ligate both fragments. | ||
We can then select the colony that does not present red color to prove that the ligation was conducted successfully. | We can then select the colony that does not present red color to prove that the ligation was conducted successfully. | ||
</p> | </p> | ||
| − | <h5 class="question">Link | + | <h5 class="question">Link <i>prk</i> with <i>ccaA</i> into pSB3K3</h5> |
| − | <p class="pcontent">We also constructed the composite part that contains both | + | <p class="pcontent">We also constructed the composite part that contains both <i>ccaA</i> and <i>prk</i>. |
| − | We construct it using the method mentioned in | + | We construct it using the method mentioned in <i>rbc</i> whole construction. |
We cloned the fragments into pSB3K3 for lower expression of PRK. | We cloned the fragments into pSB3K3 for lower expression of PRK. | ||
</p> | </p> | ||
<h5 class="question">Transformation</h5> | <h5 class="question">Transformation</h5> | ||
<p class="pcontent">After the construction of various composite parts, | <p class="pcontent">After the construction of various composite parts, | ||
| − | we co-transform them into three <i>E. coli</i> strains: BL21(DE3), W3110, and W3110(L5T7). | + | we co-transform them into three <i>E. coli</i> strains: BL21 (DE3), W3110, and W3110 (L5T7). |
| − | Since BL21(DE3) and W3110(L5T7) contains T7 polymerase, | + | Since BL21 (DE3) and W3110 (L5T7) contains T7 polymerase, |
| − | we co-transformed composite parts that | + | we co-transformed composite parts that contain T7 promoter into these strains. |
We co-transform plasmid that only contains LacI promoter into W3110. | We co-transform plasmid that only contains LacI promoter into W3110. | ||
</p> | </p> | ||
<h5 class="question">How to prove our design?</h5> | <h5 class="question">How to prove our design?</h5> | ||
<p class="pcontent">We designed a total solution test to verify the function of our whole construction. | <p class="pcontent">We designed a total solution test to verify the function of our whole construction. | ||
| − | We incubate the constructed strains in | + | We incubate the constructed strains in modified M9 medium that contains 4 (g/l) xylose as its sole carbon source. |
| − | The construction is designed to consume xylose as energy source and as a material for Calvin- | + | The construction is designed to consume xylose as energy source and as a material for Calvin-Benson cycle. |
We then measure the optical intensity (O.D. 600) to characterize the cell growth. At a fixed time interval, | We then measure the optical intensity (O.D. 600) to characterize the cell growth. At a fixed time interval, | ||
we use DNS assay to measure the sugar consumption of the bacteria. | we use DNS assay to measure the sugar consumption of the bacteria. | ||
| Line 185: | Line 222: | ||
</p> | </p> | ||
<p class="pcontent">We selected two pH sensitive promoter from <i>E. coli</i>: | <p class="pcontent">We selected two pH sensitive promoter from <i>E. coli</i>: | ||
| − | + | P<sub>asr</sub> and P<sub>gadA</sub>. P<sub>gadA</sub> will be induced under neutral condition while P<sub>asr</sub> will be induced under acidic condition. | |
| − | We cloned a sfGFP gene downstream of these promoters whose product will express green | + | We cloned a GFP and sfGFP gene downstream of these promoters respectively, whose product will express green fluorescence once the promoter has been activated. For the design of P<sub>gadA</sub> sensing system, we took the previous constructed P<sub>gadA</sub> biobrick <a href="http://parts.igem.org/Part:BBa_K1962013" |
| + | style="color:#28ff28;">BBa_K1962013</a> from <a href="https://2016.igem.org/Team:Dundee" | ||
| + | style="color:#28ff28;">2016 iGEM Dundee team</a> as our reference. We also improve the P<sub>gadA</sub> biobrick to enhance the expression of GFP. | ||
</p> | </p> | ||
<p class="pcontent">In conclusion, when the color of the medium turns from turbid yellow to green, | <p class="pcontent">In conclusion, when the color of the medium turns from turbid yellow to green, | ||
| − | it indicates the pH of the medium | + | it indicates the pH of the medium has altered so we can determine the pH condition of the medium. |
| − | + | ||
</p> | </p> | ||
<img class="gif" src="https://static.igem.org/mediawiki/2018/8/8e/T--NCKU_Tainan--design_pHsensor.gif" alt="pH"> | <img class="gif" src="https://static.igem.org/mediawiki/2018/8/8e/T--NCKU_Tainan--design_pHsensor.gif" alt="pH"> | ||
| − | < | + | <div class="row"> |
| − | <p class="pcontent">We first extracted whole genome DNA from <i>E. coli</i> MG1655 and amplify both promoters by PCR | + | <a class="btn col-md-12" data-toggle="collapse" href="#pH_how_to_construct" role="button" aria-expanded="false" aria-controls="multiCollapseExample1"> |
| − | + | How do we construct this part? | |
| − | + | <i class="fa fa-arrow-down fa-10" aria-hidden="true"></i> | |
| − | + | </a> | |
| − | + | </div> | |
| − | + | <div class="collapse multi-collapse" id="pH_how_to_construct"> | |
| − | + | <div class="card card-body"> | |
| − | + | <p class="pcontent">We first extracted whole genome DNA from <i>E. coli</i> MG1655 and amplify both promoters by PCR | |
| + | using primers that contains HindIII and SpeI. | ||
| + | We then exchanged the promoter with the previously constructed plasmid that contains P<sub>T7</sub> and GFP or sfGFP. | ||
| + | We initially transformed the constructed plasmid into DH5 alpha for colony screening. | ||
| + | We then transformed the plasmid into BL21 (DE3) to test its function. | ||
| + | We also design another biobrick that contains riboJ (a signal amplify fragment) | ||
| + | at the downstream of P<sub>gadA</sub> to get the signal more clearly and enhance the specificity. | ||
| + | </p> | ||
| + | </div> | ||
| + | </div> | ||
<img class="bigimg" src="https://static.igem.org/mediawiki/2018/d/d2/T--NCKU_Tainan--design_pH_construction.png" alt="pH alert system construction picture"> | <img class="bigimg" src="https://static.igem.org/mediawiki/2018/d/d2/T--NCKU_Tainan--design_pH_construction.png" alt="pH alert system construction picture"> | ||
| − | <h5 class="question">How we determine its function?</h5> | + | <h5 class="question">How do we determine its function?</h5> |
| − | <p class="pcontent">We measure the | + | <p class="pcontent">We measure the fluorescence intensity of the plasmid in different pH environment to |
| − | determine its promoter activity. We incubate the bacteria in pH | + | determine its promoter activity. We incubate the bacteria in pH modified M9 medium and |
| − | measure the | + | measure the fluorescence intensity (absorbance: 480 nm, excitation: 510 nm). |
</p> | </p> | ||
</div> | </div> | ||
| − | <div id=" | + | <div id="Reference"> |
<h3>References</h3> | <h3>References</h3> | ||
<ol> | <ol> | ||
| − | <li class="smallp">Cai, | + | <li class="smallp">Z. Cai, G. Liu, J. Zhang, Protein Cell (2014) 5: 552.</li> |
| − | <li class="smallp"> | + | <li class="smallp">F. Gong, “Quantitative Analysis of an Engineered CO<sub>2</sub> -Fixing <i>Escherichia Coli</i> Reveals Great Potential of Heterotrophic CO<sub>2</sub> Fixation.” Biotechnology for Biofuels, BioMed Central, 18 June 2015</li> |
| − | <li class="smallp">“The Coupling of Glycolysis and the Rubisco-Based Pathway through the Non-Oxidative Pentose Phosphate Pathway to Achieve Low Carbon Dioxide Emission Fermentation.” NeuroImage, Academic Press, 25 Mar. 2015 | + | <li class="smallp">“The Coupling of Glycolysis and the Rubisco-Based Pathway through the Non-Oxidative Pentose Phosphate Pathway to Achieve Low Carbon Dioxide Emission Fermentation.” NeuroImage, Academic Press, 25 Mar. 2015</li> |
| − | <li class="smallp">“Sugar Synthesis from CO<sub>2</sub> in <i>Escherichia Coli</i>.” NeuroImage, Academic Press, 23 June 2016 | + | <li class="smallp">“Sugar Synthesis from CO<sub>2</sub> in <i>Escherichia Coli</i>.” NeuroImage, Academic Press, 23 June 2016</li> |
| − | <li class="smallp">Cheng | + | <li class="smallp">H. Cheng, E. J. Yang, Y. L. Liu, F. Y.Chenm, Y. Ou, S. Y. Li. “The Comprehensive Profile of Fermentation Products during in Situ CO<sub>2</sub> Recycling by Rubisco-Based Engineered <i>Escherichia Coli</i>.” Microbial Cell Factories, BioMed Central, 2 Aug. 2016</li> |
</ol> | </ol> | ||
</div> | </div> | ||
| Line 236: | Line 283: | ||
} | } | ||
} else { | } else { | ||
| − | if ($(this).scrollTop() >= | + | if ($(this).scrollTop() >= 500) { |
var position = $("#sidelist").position(); | var position = $("#sidelist").position(); | ||
if(position == undefined){} | if(position == undefined){} | ||
Latest revision as of 15:18, 3 November 2018