| Line 143: | Line 143: | ||
</p> | </p> | ||
<img class="gif" src="https://static.igem.org/mediawiki/2018/3/34/T--NCKU_Tainan--design_CA.gif" alt="Rubisco"> | <img class="gif" src="https://static.igem.org/mediawiki/2018/3/34/T--NCKU_Tainan--design_CA.gif" alt="Rubisco"> | ||
| − | < | + | <div class="row"> |
| − | <p class="pcontent">We first codon optimized the sequence and insert it into the empty pSB1C3 plasmid with HindIII and | + | <a class="btn col-md-12" data-toggle="collapse" href="#CA_how_to_construct" role="button" aria-expanded="false" aria-controls="multiCollapseExample1"> |
| − | + | How do we construct this part? | |
| − | + | <i class="fa fa-arrow-down fa-10" aria-hidden="true"></i> | |
| − | + | </a> | |
| − | + | </div> | |
| + | <div class="collapse multi-collapse" id="CA_how_to_construct"> | ||
| + | <div class="card card-body"> | ||
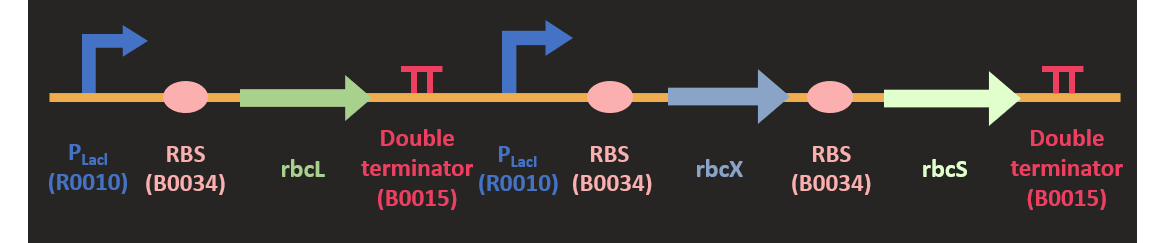
| + | <p class="pcontent">We first codon optimized the sequence and insert it into the empty pSB1C3 plasmid with HindIII and | ||
| + | SpeI just as mentioned above. In our optimized sequence, we have already designed a P<sub>T7</sub> promoter | ||
| + | in front of CA, so we can directly ligate it into the plasmid. | ||
| + | The constructed basic part is then linked with other basic parts to complete our construction. | ||
| + | </p> | ||
| + | </div> | ||
| + | </div> | ||
<img class="bigimg" src="https://static.igem.org/mediawiki/2018/7/78/T--NCKU_Tainan--design_CA_construction.png" alt="CA Construction picture"> | <img class="bigimg" src="https://static.igem.org/mediawiki/2018/7/78/T--NCKU_Tainan--design_CA_construction.png" alt="CA Construction picture"> | ||
<h5 class="question">How do we test its function?</h5> | <h5 class="question">How do we test its function?</h5> | ||
| Line 217: | Line 226: | ||
</p> | </p> | ||
<img class="gif" src="https://static.igem.org/mediawiki/2018/8/8e/T--NCKU_Tainan--design_pHsensor.gif" alt="pH"> | <img class="gif" src="https://static.igem.org/mediawiki/2018/8/8e/T--NCKU_Tainan--design_pHsensor.gif" alt="pH"> | ||
| − | < | + | <div class="row"> |
| − | <p class="pcontent">We first extracted whole genome DNA from <i>E. coli</i> MG1655 and amplify both promoters by PCR | + | <a class="btn col-md-12" data-toggle="collapse" href="#pH_how_to_construct" role="button" aria-expanded="false" aria-controls="multiCollapseExample1"> |
| − | + | How do we construct this part? | |
| − | + | <i class="fa fa-arrow-down fa-10" aria-hidden="true"></i> | |
| − | + | </a> | |
| − | + | </div> | |
| − | + | <div class="collapse multi-collapse" id="pH_how_to_construct"> | |
| − | + | <div class="card card-body"> | |
| − | + | <p class="pcontent">We first extracted whole genome DNA from <i>E. coli</i> MG1655 and amplify both promoters by PCR | |
| + | using primers that contains HindIII and SpeI. | ||
| + | We then exchanged the promoter with the previously constructed plasmid that contains P<sub>T7</sub> and GFP or sfGFP. | ||
| + | We initially transformed the constructed plasmid into DH5 alpha for colony screening. | ||
| + | We then transformed the plasmid into BL21(DE3) to test its function. | ||
| + | We also design another biobrick that contains riboJ (a signal amplify fragment) | ||
| + | at the downstream of P<sub>gadA</sub> to get the signal more clearly and enhance the specificity. | ||
| + | </p> | ||
| + | </div> | ||
| + | </div> | ||
<img class="bigimg" src="https://static.igem.org/mediawiki/2018/d/d2/T--NCKU_Tainan--design_pH_construction.png" alt="pH alert system construction picture"> | <img class="bigimg" src="https://static.igem.org/mediawiki/2018/d/d2/T--NCKU_Tainan--design_pH_construction.png" alt="pH alert system construction picture"> | ||
<h5 class="question">How do we determine its function?</h5> | <h5 class="question">How do we determine its function?</h5> | ||
Revision as of 09:40, 17 October 2018