Collaborations
USMA-West Point and USAFA Collaboration
The iGEM teams at USMA West Point and the United States Air Force Academy (USAFA) collaborated to validate the USAFA DNA submission part in the iGEM pSB1C3 plasmid. The USAFA created a bio-brick that contained an open reading frame that was intended for expression in either bacteria or yeast.
To confirm the presence of the bio-brick in the pSB1C3 plasmid, we performed PCR reactions using the intended submission part plasmid as the template, and primers (provided by the USAFA) that specifically annealed to the bio-brick insert. Prior to running the PCR reaction, the USAFA pSB1C3 plasmid was used to transform DH5alpha bacteria in order to create a bacterial stock and to generate a sufficient reserve of plasmid for all subsequent analyses.
PCR reactions were analyzed by running 10uL of 20uL PCR reactions on an 0.8% agarose gel in 1XTAE for approximately 40min at 100V. The intended iGEM submission plasmid generated a band at ~1300 bp (Figure 1, lane 2). By contrast, a control plasmid generated a band at ~1800 bp (Figure 1, lane 4). The ~1800bp product is consistent with the nucleotide sequence information provided by the USAFA team. The control plasmid used was a yeast expression plasmid into which the USAFA team previously and successfully integrated their bio-brick (pYTK-AFA18; also provided by the USAFA team). The control PCR reaction also used the primers provided by the USAFA team. The reason for the discrepancy between the PCR products for the two plasmid templates was not readily clear.
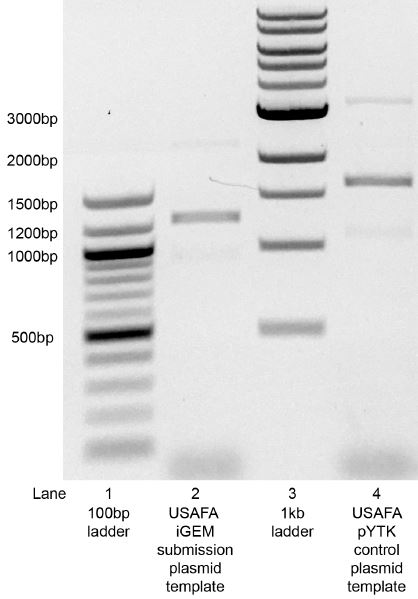
Figure 1
To independently probe the intended submission plasmid, restriction digest reactions were performed. The sequence information provided for the bio-brick insert showed that it contained two KpnI sites, whereas the iGEM pSB1C3 plasmid lacks a KpnI site (Figure 2A). If the bio-brick insert was present, then products of ~3.7 and 0.3kb were expected (Figure 2A). By contrast, two bands were observed at ~2500 and ~1500bp (Figure 2C). The second digest used EcoRI and MscI enzymes. This digest was expected to generate products 2.7, 0.7 and 0.6kb (Figure 2B). By contrast, we only observed bands at just below 1500bp and at ~700bp (Figure 2C). We believe that the ~700bp band corresponds fragment generated by the MscI and EcoRI sites present in the pSB1C3 backbone. The possibility that this apparent single band is actually the anticipated 0.6kb band unresolved from the 0.7kb band is unlikely because, as the ladder shows, products of those sizes are well resolved on this gel (restriction digest reactions were analyzed on an 0.8% agarose gel in 1XTAE for approximately 40min at 100V).
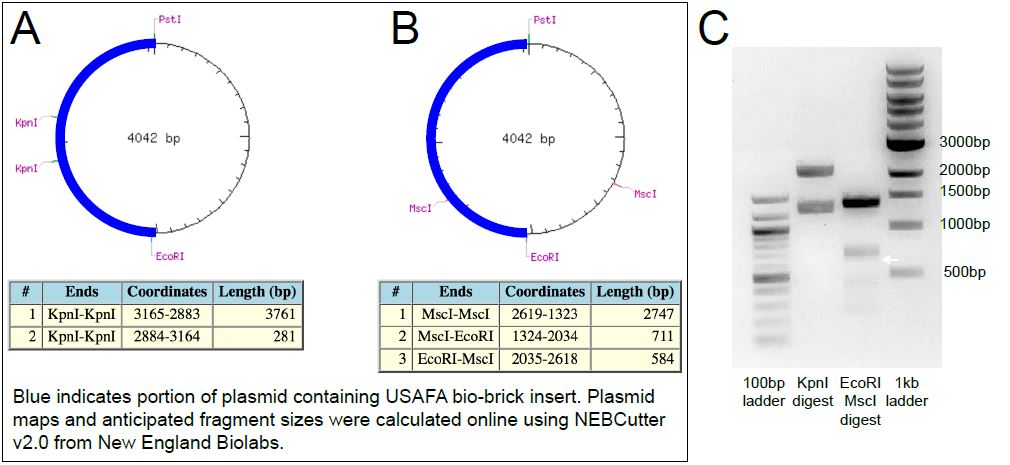
Figure 2
Due to the inconsistencies in the PCR product sizes between the reactions with the control (pYTK) and iGEM (pSB1C3) plasmids, in addition to the discrepancies in the expected and observed restriction digest products, we cannot conclude that iGEM submission plasmid is constructed as intended.


